|
Here is scientific evidence against evolution. The theory of evolution is wrong. |
Debunking Evolution
evidence that macro-evolution is impossible
|
Here is scientific evidence against evolution. The theory of evolution is wrong. |
|
Variation (microevolution) is the real part. The types of bird beaks, the colors of moths, leg sizes, etc. are variation. Each type and length of beak a finch can have is already in the gene pool and adaptive mechanisms of finches. Creationists have always agreed that there is variation within species. What evolutionists do not want you to know is that there are strict limits to variation that are never crossed, something every breeder of animals or plants is aware of. Whenever variation is pushed to extremes by selective breeding (to get the most milk from cows, sugar from beets, bristles on fruit flies, or any other characteristic), the line becomes sterile and dies out. And as one characteristic increases, others diminish. But evolutionists want you to believe that changes continue, merging gradually into new kinds of creatures. This is where the imaginary part of the theory of evolution comes in. It says that new information is added to the gene pool by mutation/natural selection to create frogs from fish, reptiles from frogs, and mammals from reptiles, to name a few.
Just to be clear, evolution theory puts no limit on what mutation/natural selection can invent, saying that everything in nature was invented by it - everything: |
|
|
|
sex, eye-hand coordination, balance, navigation systems, tongues, blood, antennae, waste removal systems, swallowing, joints, lubrication, pumps, valves, autofocus, image stabilization, sensors, camouflage, traps, ceramic teeth, light (bioluminescence), ears, tears, eyes, hands, fingernails, cartilage, bones, spinal columns, spinal cords, muscles, ligaments, tendons, livers, kidneys, thyroid glands, lungs, stomachs, vocal cords, saliva, skin, fat, lymph, body plans, growth from egg to adult, nurturing babies, aging, breathing, heartbeat, hair, hibernation, bee dancing, insect queens, spiderwebs, feathers, seashells, scales, fins, tails, legs, feet, claws, wings, beaver dams, termite mounds, bird nests, coloration, markings, decision making, speech center of the brain, visual center of the brain, hearing center of the brain, language comprehension center of the brain, sensory center of the brain, memory, creative center of the brain, object-naming center of the brain, emotional center of the brain, movement centers of the brain, center of the brain for smelling, immune systems, circulatory systems, digestive systems, endocrine systems, regulatory systems, genes, gene regulatory networks, proteins, ribosomes that assemble proteins, receptors for proteins on cells, apoptosis, hormones, neurotransmitters, circadian clocks, jet propulsion, etc. Everything in nature - according to evolution theory.
Just to be clear.
This candid admission is from the evolutionist journal Nature: "Darwin anticipated that microevolution would be a process of continuous and gradual change. The term macroevolution, by contrast, refers to the origin of new species and divisions of the taxonomic hierarchy above the species level, and also to the origin of complex adaptations, such as the vertebrate eye. Macroevolution posed a problem to Darwin because his principle of descent with modification predicts gradual transitions between small-scale adaptive changes in populations and these larger-scale phenomena, yet there is little evidence for such transitions in nature. Instead, the natural world is often characterized by gaps, or discontinuities. One type of gap relates to the existence of 'organs of extreme perfection', such as the eye, or morphological innovations, such as wings, both of which are found fully formed in present-day organisms without leaving evidence of how they evolved."-- Reznick, David N., Robert E. Ricklefs. 12 February 2009. Darwin's bridge between microevolution and macroevolution. Nature, Vol. 457, pp. 837-842
So do the big changes (macroevolution) really happen? Evolutionists tell us we cannot see evolution taking place because it happens too slowly. A human generation takes about 20 years from birth to parenthood. They say it took tens of thousands of generations to form man from a common ancestor with the ape, from populations of only hundreds or thousands. We do not have these problems with bacteria. A new generation of bacteria grows in as little as 12 minutes or up to 24 hours or more, depending on the type of bacteria and the environment, but typically 20 minutes to a few hours. There are more bacteria in the world than there are grains of sand on all of the beaches of the world (and many grains of sand are covered with bacteria). They exist in just about any environment: hot, cold, dry, wet, high pressure, low pressure, small groups, large colonies, isolated, much food, little food, much oxygen, no oxygen, in toxic chemicals, etc.
There is much variation in bacteria. There are many mutations (in fact, evolutionists say that smaller organisms have a faster mutation rate than larger ones17). But generation after generation they never turn into anything new. They always remain bacteria. Fruit flies are much more complex than already complex single-cell bacteria. Scientists like to study them because a generation (from egg to adult) takes only 9 days. In the lab, fruit flies are studied under every conceivable condition. There is much variation in fruit flies. There are many mutations. But generation after generation they never turn into anything new. They always remain fruit flies. Many years of study of countless generations of bacteria and fruit flies all over the world shows that macroevolution is not happening today.
The invention of new parts or systems by mutation has never been witnessed, nor has it been accomplished in a biochemistry laboratory. As Franklin Harold, retired professor of biochemistry and molecular biology at Colorado State University, wrote in his 2001 book "The Way of the Cell" published by Oxford University Press, "There are presently no detailed Darwinian accounts of the evolution of any biological or cellular system, only a variety of wishful speculations." Evolutionists often say "it evolved", but no one lists all the molecular steps because no one knows what they could be.
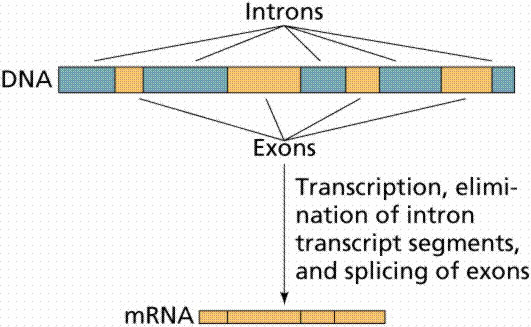
Coded information
(See
the video)
Evolutionists think
new genetic information appears randomly.
|
|
Could a million monkeys typing randomly eventually produce a copy of a play Shakespeare wrote? Maybe, but they would also fill the universe with gibberish in the process. Written languages are coded information with messages that can be intended for people or machines. Shakespeare wrote his coded information for people who read the English language code. His mind created stories to entertain, inform, and enlighten other minds. He used his hand and a pen to write the code on paper, which was transcribed onto a moveable type printing press system and published in books for other minds who understood the English code. Minds who knew different codes translated Shakespeare's information to other language codes for minds who understood them. |
The SETI Institute (search for extraterrestrial intelligence) looks for "coded information" in its search for intelligence, because intelligence and coded information go together. - https://www.seti.org/faq#obs9
Only minds make coded information and devise systems for recording and using information with meaning and purpose; everything else produces meaningless noise. It is embarrassing to have to explain the obvious, but it is necessary because evolutionists deny the obvious, yet they are trusted by most people to tell them what is true and what is not. You can trust this:
"There is no known natural law through which matter can give rise to information, neither is any physical process or material phenomenon known that can do this." Werner Gitt, page 79, 3rd English edition 2001, In the Beginning was Information, CLV, Bielefeld, Germany.
|
|
Professor of Engineering Werner Gitt specialized in information science, numerical mathematics and control engineering. He was Director and Professor at the German Federal Institute of Physics and Technology (Physikalisch-Technische Bundesanstalt Braunschweig) for over 30 years before retiring in 2003. |
And that, ladies and gentlemen, is the downfall of the Theory of Evolution. No physical process can invent the coded information needed to create any type of living thing. It is the impassable obstacle, the insurmountable wall, the unbridgeable chasm for the theory. Without coded information, natural selection has nothing to select. It doesnt matter how long you wait for something that can never happen; it will never happen. Without intelligence there is no coded information. Evolutionists talk about everything else; they talk around it; they imply that the problem has been solved; but reality is harsh; this finishes it.
| |
Some engineers write coded information for robots with instructions for making cars. DNA's information is at a higher level than any other language. It is instructions for causing the bodies of living organisms (biological machines) to form, grow, and function. Each cell has the entire genome, but it uses only the parts it needs. So if DNA is a language, who is the speaker? |

In engineering, a mind 1) recognizes a need, a useful addition, or a problem to be solved; 2) conceives a relevant idea for the purpose; 3) decides to act; 4) envisions an end product; 5) plans the required coded information, technology, and materials for assembly; 6) produces a product that serves the purpose.
Sadly, none of these are available to the Theory of Evolution. So for anything that has a purpose everything in biology you can be sure it was not made by evolution.
Evolutionists believe this: "If we look at the evolutionary record on Earth, we see a line of innovation that stretches from the origin of the first cell all the way through to our big human brains. Evolution is constantly inventing new forms and new processes in its relentless attempts to keep life going." " lets look at an innovation that has happened many times, like wings. There are many examples of evolution figuring out that wings are a useful innovation to add to a species. Insects have wings, and so do birds and bats. This tells us that the set of accidents (i.e. mutations) that leads to the emergence of wings was not very hard for evolution to stumble upon." Adam Frank. Key steps in evolution on Earth tell us how likely intelligent life is anywhere else. March 2, 2023. 13.8 blog. https://bigthink.com/13-8/key-steps-evolution-earth-intelligent-life/
|
|
Adam Frank is a self-described "evangelist of science". He is Professor of Physics and Astronomy at the University of Rochester, does research in theoretical astrophysics and plasma physics, and his research interests include astrobiology and the fluid dynamics of stars.
|
Where does Professor Frank say all that coded information came from? We are talking about billions of bits of new information between the origin of the first cell and our big human brains for changes in body plans and all the new parts and abilities. Well, it just appears like magic! He says accidents figure out useful innovations to add to species and are determined to keep life going. Sadly for evolutionists, "There is no known natural law through which matter can give rise to information, neither is any physical process or material phenomenon known that can do this." Evolutionists with their magical thinking are screwed.
|
|
How long would it take non-living physical or chemical processes to screw the screw into the hole in the wood? That's right - never; natural forces are not organized enough. It would be immensely harder for chemicals to organize into a biological machine without a DNA code guiding a living cell, or a synthetic chemist. |
Origin of
Life research
Life only comes from life. No
one has ever seen a living thing form from non-living matter, but
evolutionists are certain it happened because 1) life exists and
2) they only allow physical and chemical processes.
Evolutionists don't like to talk about "origin of life" research because it has been such a dead-end, but if chemicals never assembled themselves into the first living thing, evolution could never get started. So to keep hope alive, every once in a while over the last 70 years they have announced discoveries that supposedly bring us closer to understanding how life on Earth began.
However, the main lesson scientists have learned over those decades is that the long molecules (polymers) that allow biological creatures to work must be isolated in pure concentrations for there to be any chance of success. Unfortunately, that can only happen in biochemistry labs, computer simulations, and living cells. In all other settings, the products are unusable due to contamination, unwanted reactions with other chemicals, and minuscule concentrations that quickly fall apart.
|
|
Amino acids are often called the "building blocks of life". Most people know of an "experiment published in 1953 by Stanley Miller. He applied a spark discharge to a mixture of simple gases that were then thought to represent the atmosphere of the early Earth. Two amino acids of the set of 20 used to construct proteins were formed in significant quantities, with others from that set present in small amounts." - Shapiro, Robert. June 2007. A Simpler Origin for Life. Scientific American, Vol. 296, pp. 24-31. |
That was over 70 years ago. Efforts to build a living cell from scratch
in a lab have gone nowhere, so in 2017 the Build-A-Cell
project was launched to allow synthetic chemists everywhere to work together
and maybe make some progress. This is from their
Build-A-Cell website: "Cells are the fundamental 'building blocks' that make up
living organisms. Yet, we don't know exactly how cells were formed in the first
place. We also don't know what all the molecules that make up any natural cell
do. Finally, we can't yet put molecules
together ourselves to make new synthetic cells."
https://www.buildacell.org/#:~:text=Build%2DA%2DCell%20is%20an,a%20diversity%20of%20synthetic%20cells.
They don't know exactly how cells were formed in the first place? In fact, they have no clue.
Synthetic organic chemist par excellence Dr. James Tour offered a suggestion in a 2023 interview:
|
|
"A resurrection should be easier than a bottom-up synthesis. "If I gave you a cell that just died; go ahead bring it back to life." "Were just talking about a little cell, a yeast cell, a very simple cell, not even human cells." "Everythings there; all the parts are in
place. Now bring it back to
life. Can anybody do that? There is not a scientist in their right
mind who will say that they can do that.
Even origin of life people would never say that they can do that. They wont say they cant do it, because they wont admit it. Theyll just look at you. Thats what they do, they just look at you." "I saw Steve Benner (a big origin of life
researcher)
I challenged him on this.
No - he just stared at me; they cant do it." |
|
|
An interview with Steven A. Benner,
Ph.D. Chemistry, Harvard, prominent origin-of-life researcher and
creator of the Foundation for Applied Molecular Evolution, was posted
on Huffington Post on December 6, 2013. In it he said, "We
have failed in any continuous way to provide a recipe that gets
from the simple molecules that we know were present on early Earth
to RNA." "The first paradox is the tendency of organic
matter to devolve and to give tar. If you can avoid that,
you can start to try to assemble things that are not tarry, but
then you encounter the water problem, which is related to the fact
that every interesting bond that you want to make is unstable, thermodynamically,
with respect to water. If you can solve that problem, you
have the problem of entropy, that any of the building blocks are
going to be present in a low concentration; therefore, to assemble
a large number of those building blocks, you get a gene-like RNA
-- 100 nucleotides long -- that fights entropy. And the fourth
problem is that even if you can solve the entropy problem, you have
a paradox that RNA enzymes, which are maybe catalytically active,
are more likely to be active in the sense that destroys RNA rather
than creates RNA." |
Two prominent "origin-of-life" researchers have laid out their vision of how life arose from chemicals: (see my video on this)
|
|
|
1. Start with a molecule capable of copying itself. "The first protocells contained RNA (or something similar to it) and little else". 2. A fatty acid bubble forms around the self-copying molecule, which then makes a copy of itself with nucleotides that filter through the bubble. "Molecules as large as nucleotides can in fact easily slip across membranes as long as both nucleotides and membranes are simpler, more 'primitive' versions of their modern counterparts." |
3. The double-strand RNA separates into single strands if it is heated just right. That might happen in an icy pond next to a volcano, where the bubble could circulate between the ice and the hot rocks. "The sudden heating would cause a double helix to separate into single strands. Once back in the cool region, new double strands, copies of the original, could form". At the same time, the bubble is picking up fatty acid molecules and growing. Adding fatty acids makes the membrane grow longer, and a little shaking breaks the bubble into some smaller bubbles, each with some of the self-copying molecules inside, so you have "cell division".
4. "At some point some of the RNA sequences mutated, becoming ribozymes". The "ribozymes (folded RNA molecules analogous to protein-based enzymes) arise and take on such jobs as speeding up reproduction and strengthening the protocell's membrane. Consequently, protocells begin to reproduce on their own." "Other ribozymes catalyze metabolism -- chains of chemical reactions that enable protocells to tap into nutrients from the environment."
5. "Next, the organisms might have added protein-making to their bag of chemical tricks." "Complex systems of RNA catalysts begin to translate strings of RNA letters (genes) into chains of amino acids (proteins)." "Proteins take on a wide range of tasks within the cell."
6. "Protein-based catalysts, or enzymes, gradually replace most ribozymes." "Proteins would have then taken over RNA's role in assisting genetic copying and metabolism."
7. Later, the organisms would have 'learned' to make DNA". "Thanks to its superior stability, DNA takes on the role of primary genetic molecule. RNA's main role is now to act as a bridge between DNA and proteins."
8. "Organisms resembling modern bacteria adapt to living virtually everywhere on earth and rule unopposed for billions of years, until some of them begin to evolve into more complex organisms."-- Ricardo, Alonso, Jack W. Szostak. September 2009. The Origin of Life on Earth. Scientific American, pp. 54-61.
They are currently working on steps 1 and 2 in the laboratory.
Let's compare their "origin of life" ideas to the plans of children making a spaceship out of a cardboard box:
|
|
|
1. Get a large box. Draw controls and gauges on the inside. Cut out a door and round windows. Attach cardboard fins to the sides. 2. Put a chair in the box, sit down and start the countdown. |
3. Launch the spaceship towards the Moon. Using the Moon's gravity, fling the spaceship to the outer reaches of the solar system, constantly accelerating with the impulse engines.
4. After passing Neptune, engage the warp drive in a direction perpendicular to the plane of the ecliptic to avoid the Kuiper belt.
The children are currently working on steps 1 and 2, and are as close to fulfilling their goal as the "origin-of-life" researchers are.
 Franklin M. Harold studied cell biology for over
50 years. Researcher William F. Martin called him "a grand master of cellular
workings and bioenergetics" in a BioEssays book review. Harold Is
Professor Emeritus, Department of Biochemistry and Molecular Biology, Colorado
State University, Fort Collins, Colorado, and Affiliate Professor, Department
of Microbiology, University of Washington Health Sciences Center, Seattle,
Washington. In a chapter titled
"Ultimate Riddle - Origin of Cellular Life" in his 2014 book "In Search of Cell
History: The Evolution of Lifes Building Blocks" published by the University
of Chicago Press, he examined at length the
current state of origin-of-life research.
These are some of his conclusions:
Franklin M. Harold studied cell biology for over
50 years. Researcher William F. Martin called him "a grand master of cellular
workings and bioenergetics" in a BioEssays book review. Harold Is
Professor Emeritus, Department of Biochemistry and Molecular Biology, Colorado
State University, Fort Collins, Colorado, and Affiliate Professor, Department
of Microbiology, University of Washington Health Sciences Center, Seattle,
Washington. In a chapter titled
"Ultimate Riddle - Origin of Cellular Life" in his 2014 book "In Search of Cell
History: The Evolution of Lifes Building Blocks" published by the University
of Chicago Press, he examined at length the
current state of origin-of-life research.
These are some of his conclusions:
Over the past sixty years, dedicated and skillful scientists have devoted much effort and ink to the origin of life, with remarkably little to show for it. [Quoting Radu Popa, 2004,] "So far, no theory, no approach, no set of formulas, and no blackboard scheme has been found satisfactory in explaining the origin of life." At the conclusion of a century of science, whose great glory is the discovery of how living things work, there is something downright disgraceful about this confession, an intimation that despite our vast knowledge and clever technology there may be questions that exceed our grasp. But its truth is indisputable. A survey of the literature devoted to the beginnings of life leaves one in no doubt that all the critical questions remain open. For the present, we are in limbo. The natural path from simple cosmic molecules to cells, from chemistry to biology, remains undiscovered. where we should look for illumination I cannot say. The difference between a puzzle and a mystery is that the former can be solved within the framework of known principles, while the latter cannot. In the end, the origin of life remains a mystery that passes understanding. we may still be missing some essential insight. Scientists' refusal to grant some space to the mind and will of God may strike the majority of mankind as arbitrary and narrow-minded, but it is essential if the origin of life is to remain within the domain of science. A nudge from the divine would help us clear some very high hurdles; but once that possibility is admitted there will be no place to stop, and soon the settled principle of evolution by natural selection would be thrown into doubt. Life's origin has been most ardently pursued by chemists, apparently on the unspoken premise that once the molecular building blocks are on hand, cellular organization will take care of itself. That premise is surely incorrect. Modern cells do not assemble themselves from preformed constituents, and they would not have done so in the past. the notion that the first protocells assembled themselves spontaneously from a generous menu of precursor molecules conveniently supplied by abiotic chemistry (or imported by way of comets and meteorites) is now widely recognized as simplistic and effectively has been abandoned. Among its most cogent critics are experienced masters of the art of prebiotic synthesis, who are well aware of the shortcomings of many of the proposed routes and of the wide gap between the range of molecules that living things employ and those that can be made in the laboratory. the fact is that chemists have encountered insuperable difficulties in generating a working replicator, and many have expressed doubts about the project. It is at least incumbent upon proponents of its spontaneous genesis to explain how the "correct" monomers could have been selected from the "prebiotic clutter," how a sufficient concentration of monomers was maintained, where the energy came from, and how the replicator evaded the tendency of polymers to break down by hydrolysis. A decade ago, a hot topic for debate was which came first, replication or metabolism? That issue has not been resolved but has been largely superseded by the recognition that neither of them, by itself, can take one far along the road to life. It is simply not credible to claim that anything beyond the most rudimentary kind of replication or metabolism could have arisen in free solution. In truth, there is presently no persuasive hypothesis to account for the emergence of protocells from the primal chaos. The crucial step in the transfiguration of protocells into true cells will have been the invention of translation and the genetic code. the origin of the principles that govern cellular operations today - genes specifying proteins and all the apparatus that this requires - remains quite unknown and points beyond the capacity of present-day biochemistry and biophysics. |
|
|
If you want to know about origin of life research from an expert in synthetic organic chemistry, you can't do any better than Dr. James M. Tour, Ph.D. in synthetic organic and organometallic chemistry from Purdue with postdoctoral training in synthetic organic chemistry at the University of Wisconsin and Stanford, currently Professor of Chemistry, Computer Science, Materials Science and NanoEngineering at Rice University. |
In a 2016 lecture at https://www.youtube.com/watch?v=_zQXgJ-dXM4 Dr. James M. Tour said, "From a synthetic chemical perspective, neither I nor any of my colleagues can fathom a prebiotic molecular route to construction of a complex system. We cannot even figure out the prebiotic routes to the basic building blocks of life: carbohydrates, nucleic acids, lipids, and proteins. Chemists are collectively bewildered. Hence I say that no chemist understands prebiotic synthesis of the requisite building blocks, let alone assembly into a complex system. That's how clueless we are. I have asked all of my colleagues - National Academy members, Nobel Prize winners - I sit with them in offices. Nobody understands this. So if your professors say it's all worked out, if your teachers say it's all worked out, they don't know what they're talking about." "We have no idea how the molecules that compose living systems could have been devised such that they would work in concert to fulfill biology's functions... Those that say, 'Oh this is well worked out', they know nothing - nothing - about chemical synthesis - nothing."
He answers the most important questions about origin of life from chemicals in this video at https://www.youtube.com/watch?v=r4sP1E1Jd_Y
Read his 2017 "An Open Letter to My Colleagues" at http://inference-review.com/article/an-open-letter-to-my-colleagues
And so the faithful Origin of Life researchers continue to strive, undaunted, year after year, decade after decade, to solve the impossible mystery. As long as evolutionists can say that scientists are working on it, the rest of the world doesnt notice that the research is stalled where it began.
The
smallest living genome
How
many genes would it take to make Evolution's first living cell in that
legendary primordial soup? Unfortunately
for Origin of Life researchers, the smallest self-duplicating living cell found
in nature doesn't have just one or two genes; it is Mycoplasma genitalium, with 525 genes. But in 2016 Craig Venters lab got the Mycoplasma mycoides genome down to 473 genes in 531,000 base pairs. They were "interested in simplifying the
genomic software of a bacterial cell by eliminating genes that are nonessential
for cell growth under ideal conditions in the laboratory" in order to gain an "understanding
of the molecular and biological function of every gene that is essential for life." Here is what those genes do:
Hutchison III, Clyde A. et al. 25 March 2016. Design and synthesis of a minimal bacterial genome. Science, Vol. 351, No. 6280, aad6253-1-11 DOI:10.1126/science.aad6253
|
|
Nipped in the bud |
The bacteria, archaea, and single-celled eukaryotes at the very base of this "tree" would have used and expanded the mechanisms of the first living cell through "descent with modification", Darwins definition of evolution.
From: Spang, Anja, Thijs J.G. Ettema. 26 April 2016. Microbial diversity: The tree of life comes of age. Nature Microbiology, Vol. 1, No. 5, 16056 DOI:10.1038/nmicrobiol.2016.56
Reality, however, turns out to be different:
"[E]ach of the three domains of life (bacteria, archaea, and eukaryotes) has its own unique way of 1) replicating its genome, 2) defining whether a piece of DNA is a gene or not, 3) determining whether an RNA is protein-coding or not, and 4) if it is protein-coding, where transcription or translation should start and end."
"To borrow the language of cryptography, the inheritable genetic information that determines life or death of all living beings is encrypted. Furthermore, the information is encrypted in different ways in different organisms, one way for bacteria, another way for archaea, and yet another way for eukaryotes. These organisms have to use their own unique cryptographic keys to decipher their genomes. Their cryptographic keys are their own RNAs and proteins present in their own cells and those that they can make themselves using their own molecular machineries."
"It is like Chinese and English - they use totally different alphabets, words, and grammars and need to be read differently."
"[T]he same task is implemented differently by the three fundamental cell types. That is not what one would expect if bacteria and eukaryotes had shared a common ancestor because DNA replication is essential for the survival and reproduction of each and every known organism."
"This creates unbridgeable gaps between bacteria, archaea, and eukaryotes and, thus, challenges the popular belief that life came from non-life naturally and that all organisms are connected via a big evolutionary tree of life."
"What many believe and teach about the origin of life and
the origin of biodiversity does not agree with what the genes are showing us."
- Tan, Change Laura.
2022. Facts Cannot be Ignored When Considering the Origin of Life #3: Necessity
of Matching the Coding and the Decoding Systems. Answers Research Journal, Vol.
15, pp. 4960.
DOI:assets.answersresearchjournal.org/doc/v15/origin_of_life_coding_decoding.pdf
| |
Associate Professor Change Laura Tan teaches molecular biology at the University of Missouri. Her Ph.D. is from the University of Pennsylvania in biochemistry (developmental biology), with postdoctoral work in genetics at Harvard Medical School. |
Like most of us, Dr. Tan believed what she was taught about evolution, until her work led her to see problems with the theory. Fearlessly publishing her discoveries that undermine Evolution theory in a creationist peer-reviewed journal (Answers Research Journal) and a book led to her losing tenure at the university. Evolutionists are fond of repeating the story of the persecution of Galileo by the Church in 1633 as evidence of ignorant, bigoted thinking, but they are guilty of it themselves many times over. Evolutionists try to keep their oppression quiet, but feel their punitive actions are justified because they are working for a higher cause: protecting the truth from heresy, same as the Taliban. Such retribution is regrettably common. - https://crev.info/2023/01/tenure-no-longer-protects-creationist-professors/
Mutation -
natural selection
Here is how the imaginary part
is supposed
to happen: On rare
occasions a mutation in DNA improves a creature's
ability to survive, so it is more likely to reproduce (natural
selection). That is evolution's only tool for making new creatures.
It might even work if it took just one
gene to make and control one part. But
parts of living creatures are constructed of intricate
components with connections that all need to be in place for the thing to
work, controlled by many genes that have to act in the proper sequence. Natural selection would not choose parts that did
not have all their components existing, in place, connected, and
regulated because the parts would not work. Thus all
the right mutations (and none of the destructive ones) must be present
at the same time by pure chance. That is physically impossible.
To illustrate just how hopeless it is, imagine this: on the ground are all the materials needed to build a house (nails, boards, shingles, windows, etc.). We tie a hammer to the wagging tail of a dog and let him wander about the work site for as long as you please, even millions of years. The swinging hammer on the dog is as likely to build a house as mutation-natural selection is to make a single new working part in an animal, let alone a new creature.

Only mutations in the reproductive (germ) cells of an animal or plant would be passed on. Mutations in the eye or skin of an animal would not matter. Mutations in DNA happen fairly often, but most are repaired or destroyed by mechanisms in animals and plants. All known mutations in animal and plant germ cells are neutral, harmful, or fatal. But evolutionists are eternally optimistic. They believe that millions of beneficial mutations built every type of creature that ever existed.
Believing in beneficial mutations is like believing a short-circuit in the motherboard of your computer could improve its performance. To make any lasting change, a beneficial mutation would have to spread ("sweep") through a population and stay (become "fixed"). To evolutionists, this idea has been essential for so long that it is called a "classic sweep", "in which a new, strongly beneficial mutation increases in frequency to fixation in the population."
Some evolutionist researchers went looking for classic sweeps in humans, and reported their findings in the journal Science. "To evaluate the importance of classic sweeps in shaping human diversity, we analyzed resequencing data for 179 human genomes from four populations". "In humans, the effects of sweeps are expected to persist for approximately 10,000 generations or about 250,000 years." Evolutionists had identified "more than 2000 genes as potential targets of positive selection in the human genome", and they expected that "diversity patterns in about 10% of the human genome have been affected by linkage to recent sweeps." So what did they find? "In contrast to expectation," their test detected nothing, but they could not quite bring themselves to say it. They said there was a "paucity of classic sweeps revealed by our findings". Sweeps "were too infrequent within the past 250,000 years to have had discernible effects on genomic diversity." "Classic sweeps were not a dominant mode of human adaptation over the past 250,000 years." --Hernandez, Ryan D., Joanna L. Kelley, Eyal Elyashiv, S. Cord Melton, Adam Auton, Gilean McVean, 1000 Genomes Project, Guy Sella, Molly Przeworski. 18 February 2011. Classic Selective Sweeps Were Rare in Recent Human Evolution. Science, Vol. 331, no. 6019, pp. 920-924.
|
|
A 35-year experiment by evolutionists shows how things really work. Instead of waiting for natural selection, researchers forced selection on hundreds of generations of fruit flies. They used variation to breed fruit flies that develop from egg to adult 20% faster than normal. But, as usual when breeding plants and animals, there was a down side. In this case the fruit flies weighed less, lived shorter lives, and were less resistant to starvation. There were many mutations, but none caught on, and the experiment ran into the limits of variation. |
They wrote that "forward experimental evolution can often be completely reversed with these populations". "Despite decades of sustained selection in relatively small, sexually reproducing laboratory populations, selection did not lead to the fixation of newly arising unconditionally advantageous alleles." "The probability of fixation in wild populations should be even lower than its likelihood in these experiments." --Burke, Molly K., Joseph P. Dunham, Parvin Shahrestani, Kevin R. Thornton, Michael R. Rose, Anthony D. Long. 30 September 2010. Genome-wide analysis of a long-term evolution experiment with Drosophila. Nature, Vol. 467, pp. 587-590.
|
|
You may have heard of the famous Lenski experiment. Dr. Richard E. Lenski is an evolutionary biologist who began a long-term experiment on February 24, 1988 that continued till May 2022 at Michigan State University. It looked for genetic changes in 12 initially identical populations of Escherichia coli bacteria that adapted to conditions in their flasks for 75,000 generations. |
I have simplified a report by Scott Whynot, who studied 26 peer-reviewed scientific articles authored by Dr. Lenski (with others) published between 1991 and 2012. These papers represent the major genetic findings from 21 years of the experiment.
1. There was an insertion mutation that inhibited transcription of DNA involved in cell wall synthesis.
2. There was an insertion mutation in a regulatory region that encodes two proteins involved with cell wall synthesis. This may have led to larger cells.
3. A mutation in a gene led to a defect in DNA repair.
4. An insertion mutation may have knocked out a gene involved in programmed cell death and response to stress.
5. There was another mutation in a gene involved in response to stress, disrupting its function.
6. There was a mutation in the gene that encodes an enzyme that loosens DNA coils, leading to an increase in DNA supercoiling.
7. There was an insertion mutation in a gene that represses the production of nicotinamide adenine dinucleotide (NAD), a molecule that participates in many metabolic reactions, some affecting longevity. This might allow more NAD production.
8. The researchers noted an insertion mutation that they think inactivated a gene, resulting in greater glucose uptake. Glucose is a limited energy source in the experiment.
9. Deletion mutations caused the loss of the ability to catabolize D-ribose, an energy source that is not available in the experiment.
10. There was a mutation in a gene regulating transport of the sugar maltose, an energy source that is not present in the experiment.
11. After about 30,000 generations, the E. coli in one of the twelve isolated populations began to utilize an energy source, citrate, that they normally could not use in the presence of oxygen. E. coli already have the ability to transport and metabolize citrate where there is no oxygen, but they do not produce an appropriate transport protein for an environment with oxygen. In E. coli DNA, the gene for the citrate transporter that works without oxygen is directly upstream from genes for proteins with promoters that are active in the presence of oxygen. A replication of the region happened to put the transporter gene next to one of these promoters, so it could now be expressed in the presence of oxygen.
Except for number 11, the changes found in over 60,000 generations of bacteria were due to the disruption, degradation, or loss of genetic information. The ability to use citrate in the presence of oxygen, trumpeted by evolutionists as a big deal, was the result of previously existing information being rearranged, not the origin of new information. Mutations that result in a gain of novel information have not been observed.
"Most long-term evolution experiments thus far have been performed in bacteria or haploid yeast populations, where, in most environments, there exist a number of loss-of-function mutations that provide a selective advantage." "For instance, sterility in yeast provides a selective advantage by eliminating unnecessary gene expression." "The emergence of the Cit+ phenotype is the exception in experimental evolution, where most evolved mutations affect independent genes and biological pathways, driven largely by large-target loss-of-function mutations."-- Lang, Gregory I., Michael M. Desai. 2014. The spectrum of adaptive mutations in experimental evolution. Genomics, Vol. 104, No. 6, Part A, pp. 412416.
Imagine the coded information required to grow an animal from an egg cell (ontogeny). This is what researchers have discovered about animal body plans:
"The overall control principle is that the embryonic process is finely divided into precise little 'jobs' to be done, and each is assigned to a specific subcircuit or wiring feature in the upper level dGRN [developmental gene regulatory network]. No subcircuit functions are redundant with another, and that is why there is always an observable consequence if a dGRN subcircuit is interrupted. Since these consequences are always catastrophically bad, flexibility is minimal, and since the subcircuits are all interconnected, the whole network partakes of the quality that there is only one way for things to work. And indeed the embryos of each species develop in only one way."
There is no place for mutation-natural selection here.
" mechanistic developmental biology has shown that its fundamental concepts are largely irrelevant to the process by which the body plan is formed in ontogeny [a developing embryo]. In addition it gives rise to lethal errors in respect to evolutionary process. NeoDarwinian evolution is uniformitarian in that it assumes that all process works the same way, so that evolution of enzymes or flower colors can be used as current 4 proxies for study of evolution of the body plan. It erroneously assumes that change in protein coding sequence is the basic cause of change in developmental program; and it erroneously assumes that evolutionary change in body plan morphology occurs by a continuous process. All of these assumptions are basically counterfactual. This cannot be surprising, since the neo-Darwinian synthesis from which these ideas stem was a pre-molecular biology concoction focused on population genetics and adaptation natural history, neither of which have any direct mechanistic import for the genomic regulatory systems that drive embryonic development of the body plan."
"No observations on single genes can ever illuminate the overall mechanisms of the development of the body plan or of body parts".-- Davidson, Eric H. 1 September 2011. Evolutionary bioscience as regulatory systems biology. Developmental Biology, Vol. 357, No. 1, pp. 35-40 DOI:10.1016/j.ydbio.2011.02.004



Waiting for mutations
Evolutionists believe that humans share a common ancestor with the
great apes of Africa. They say "hominins"
are the human lineage arising from that ancestor. A 2015
paper calculated how long it would take to change the nucleotides in hominin
DNA. These excerpts from it will shock you:
"Given the unique capabilities of humans, an evolving hominin population (as would give rise to modern man) would need to establish a great deal of new information."
"It is estimated that it only took six million years for the chimp and human genomes to diverge by over 5%, representing about 150 million nucleotide differences."
"The gene can range in size from about 1,000 to more than one million nucleotides long. A typical human gene is roughly 50,000 nucleotides long. A new gene is thought to arise from a previously existing gene, with the mutation/selection process establishing mutations within a long text string that is already established and functional."
"It is now generally recognized that beneficial mutations are rare, and that high-impact beneficial mutations are extremely rare. In higher life forms where population sizes are modest, the mutation rate per nucleotide per generation is normally extremely low (about 10−8). This means that the waiting time for a specific nucleotide within single chromosomal lineage would be 100 million generations."
"We simulated a classic pre-human hominin population of at least 10,000 individuals, with a generation time of 20 years, using the numerical simulation program Mendels Accountant (Mendel version 2.4.2, now being released as 2.5)."
"Biologically realistic numerical simulations revealed that a population of this type required inordinately long waiting times to establish even the shortest nucleotide strings. To establish a string of two nucleotides required on average 84 million years. To establish a string of five nucleotides required on average 2 billion years. We found that waiting times were reduced by higher mutation rates, stronger fitness benefits, and larger population sizes. However, even using the most generous feasible parameter settings, the waiting time required to establish any specific nucleotide string within this type of population was consistently prohibitive."
"Even given very substantial fitness effects, the waiting time for a specific point mutation ranged between 1.5 and 15.9 million years" which "is very sobering, since it is estimated that mankind evolved from a chimp-like creature in just 6 million years."
"As string length increased linearly, the increase in waiting time was of an exponential nature. When there were as many as six nucleotides in the string, the average waiting time (4.24 billion years) approached the estimated age of the earth. When there were eight nucleotides in the string, the average waiting time (18.5 billion years), exceeded the estimated age of the universe."
"Our results generally represent best-case scenarios in terms of minimizing waiting time. When we use more realistic parameter settings for our simulations, we consistently get much longer waiting times."
"When a population faces a specific evolutionary challenge, a specific fix is needed, and it must arise in a timely fashion. Positive selection cannot generally begin to resolve an evolutionary challenge until just the right mutation (or mutations) happens at just the right position (or positions). Selection for the required trait can only begin after the mutation (or mutations) result in a substantial (selectable) improvement in total biological functionality."
"The creation and fixation of a string of three (requiring at least 380 million years) would be extremely untimely adaptation in the face of any type of pressing evolutionary challenge (and trivial in effect), in terms of the evolution of modern man" who has "a genome with over three billion nucleotides."
"We need multiple point mutations to arise on the same short strand of DNA, which is very difficult. While a population is waiting (through deep time) for the correct string to arise, genetic drift is systematically eliminating almost all the string variants. Nearly all of the time there will be essentially zero strings anywhere in the population that are even close to the target string."
"It is widely thought that a larger population size can eliminate the waiting time problem. While our simulations show that larger populations do help reduce waiting time, we see that the benefit of larger population size produces rapidly diminishing returns. When we increase the hominin population from 10,000 to 1 million, the waiting time for creating a string of five is only reduced from two billion to 482 million years. This amount of time approximates the estimated time required for the evolution of worm-like creatures into people. When we extrapolate our data to a population size of ten million we still get a waiting time of 202 million years. Even when we extrapolate to a population size of one billion we still have a waiting time of 40 million years."
"A bigger population increases the number of mutations arising per generation, but does not increase the number of mutations per short DNA strand (mutation density). To create a complete set of linked mutations requires many mutations arising on the same short stretch of a given DNA molecule."
"Numerous other researchers have come to similar conclusions. The long waiting times we report here are even supported indirectly by the papers that have argued against a serious waiting time problem. When examined carefully, those papers indicate that for a hominin-type population, waiting times are as long or even longer than we report here."
It is true that "during the waiting time period for a functional string to be established at a given location, other beneficial mutational strings can be happening in other parts of the genome."
"However, those other strings are not likely to meet the same specific evolutionary need that our target string can meet. Evolution often needs a specific fix to a specific problem, and that fix must be timely in order to retain relevance."
"Even if all of the ~20,000 genes in the hominin genome were already poised for a significant enhancement and all of them were waiting for their own specific string, each one of those potential enhancements would have its own severe waiting time problem."
"Furthermore, this would be happening in the context of countless nearly-neutral deleterious mutations throughout the genome which would drift to fixation within the same deep time. Unless there was very strong purifying selection operating for all the nucleotides in the general region of the string, the context of the string would be erased long before the string itself actually arose."
"The waiting time problem becomes very severe when more than one mutation is required to establish a new function. This is a very interesting theoretical dilemma."-- Sanford, John, Wesley Brewer, Franzine Smith and John Baumgardner. September 17, 2015. The waiting time problem in a model hominin population. Theoretical Biology and Medical Modelling, Vol. 12, No. 1, Article 18, 28 pages, DOI: 10.1186/s12976-015-0016-z.
Orphan genes
- the final blow?
|
|
|
Here is an evolutionist with experience in molecular biology, Francois Jacob. Francois Jacob won the Nobel Prize in Physiology or Medicine in 1965, along with two others, for discoveries concerning genetic control of enzyme and virus synthesis. He had joined the Institut Pasteur in 1950. He was appointed Laboratory Director there in 1956, then Head of the Department of Cell Genetics in 1960. In 1964 he was appointed Professor at the College de France, where a chair of Cell Genetics was created for him. He was Chairman of the Board of the Institut Pasteur from 1982 to 1988. The work of Francois Jacob dealt mainly with the genetic mechanisms existing in bacteria and bacteriophages, and with the biochemical effects of mutations. |
He wrote, "Evolution does not produce novelties from scratch. It works on what already exists, either transforming a system to give it new functions or combining several systems to produce a more elaborate one."
"During chemical evolution in prebiotic times and at the beginning of biological evolution, all those molecules of which every living being is built had to appear. But once life had started in the form of some primitive self-reproducing organism, further evolution had to proceed mainly through alterations of already existing compounds. New functions developed as new proteins appeared. But these were merely variations on previous themes. A sequence of a thousand nucleotides codes for a medium-sized protein. The probability that a functional protein would appear de novo by random association of amino acids is practically zero. In organisms as complex and integrated as those that were already living a long time ago, creation of entirely new nucleotide sequences could not be of any importance in the production of new information."20
For decades, everyone agreed. But as researchers compared the genes of similar creatures, they found that the genes differed, from just a little to a lot. They imagined different ways that could have happened. Gene duplication, non-deleterious frame shift mutations, alternative reading frames, overlap with transposable elements, horizontal gene transfer, or overlapping gene.45 As usual with evolutionists, they do not know what really happened, they assume it was one of these mental explanations, and that is enough. But some genes are so unique, even imagination fails. Evolutionists now conclude they must have assembled spontaneously - "de novo". In fact, "all genome and expressed sequence tag (EST) projects to date in every taxonomic group studied so far have uncovered a substantial fraction of genes that are without known homologs [equivalents]. These 'orphans' or 'taxonomically restricted genes' (TRGs) are defined as being exclusively restricted to a particular taxonomic group."21 "Orphan genes are defined as genes which lack detectable similarity to genes in other species". "They typically make up 10 to 30% of all genes in a genome."45
The foundation of evolution theory, gradual modification over time, slowly transforming genes that already exist, suddenly ran up against orphan genes, genes without parents in every taxonomic group studied so far. Looking at it objectively, the theory of evolution has been falsified. After careful study, evolutionists made a bold choice:
|
|
They cut the theory's last connection to reality, declaring that the impossible is normal: of course genes are produced de novo! The new foundation of evolution theory is Poof - there it is (which sounds like the foundation of creation by Intelligent Design - de novo).
Evolutionists now think orphan genes are awesome. "There should be greater appreciation of the importance of the de novo origination of genes." "Today, we know that this evolutionary process is not impossible."47 "De novo evolution is clearly a strong force - constantly generating new genes over time." "It seems possible that most orphan genes have evolved through de novo evolution."35 "It looks as if we couldn't find the families of most orphans because they don't really have families."35 "The sequencing of a large number of eukaryotic and bacterial genomes has uncovered an abundance of genes without homologs... and has shown that new genes have arisen in the genomes of every group of organisms studied so far including humans".21 |
For evolutionists, the theory of evolution can never die. The rest of us can see that Francois Jacob was right. Orphan genes reveal that macro-evolution does not represent reality, and is physically impossible.
Genes are hundreds to thousands of base pairs long. How many orphan genes are in animals? Here are some examples (numbers are approximate based on available data):
Saccharomyces cerevisiae (budding yeast) has 1192 orphan
genes
Schizosaccharomyces pombe (fission yeast)
has 691 orphan genes
Caenorhabditis elegans (the worm)
has 2200 orphan genes
Pristionchus pacificus (parasitic
nematode) has 7050 orphan genes
Ciona intestinalis (sea vase or vase
tunicate) has 3200 orphan genes
Drosophila melanogaster (common fruit
fly) has 2604 orphan genes
Drosophila simulans (fruit fly) has 2937 orphan genes
Drosophila erecta (fruit fly in West
Africa) has 2071 orphan genes
Drosophila pseudoobscura (fruit fly in
western North America) has 3206 orphan
genes
Drosophila persimilis (fruit fly)
has 4362 orphan genes
Aedes aegypti (yellow fever mosquito)
has 3956 orphan genes
Tribolium castaneum (red flour beetle)
has 3200 orphan genes
Mus musculus (house mouse) has 243 orphan genes
Rattus norvegicus (Norway rat) has 2513 orphan genes
Ornithorhynchus anatinus (duck-billed
platypus) has 3330 orphan genes
Bos Taurus (domesticated cattle) has
2384 orphan genes
Homo sapiens (humans) have 1398 orphan genes
Percentages of orphan genes from: Khalturin, Konstantin, et al. September 2009. More than just orphans: are taxonomically-restricted genes important in evolution? Trends in Genetics, Vol. 25, No. 9, pp. 404-413 DOI:10.1016/j.tig.2009.07.006
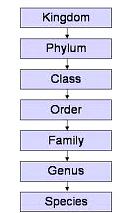
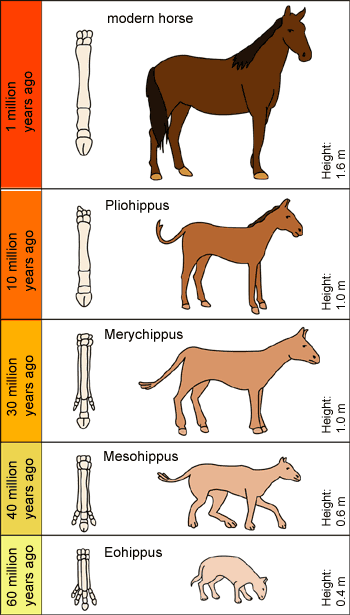
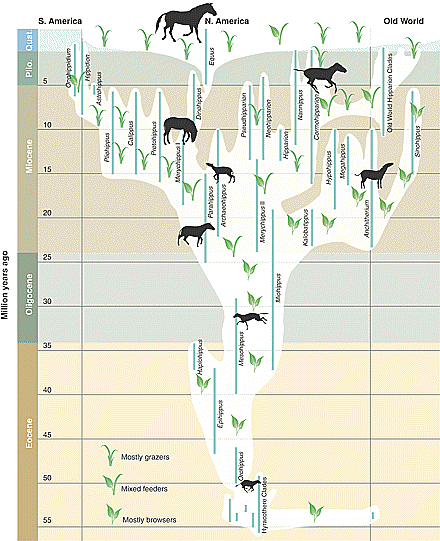
No more lines
When you want to sell an idea to people start
with something they are familiar with, like a family tree.
Evolutionists did that with macro-evolution, and people quickly fell for it. Whats the difference? The members on this tree are not related because macroevolution is physically impossible, as this article proves.
Without a source for coded information, a path to the origin of life, rapid production of beneficial mutations, and an excuse for orphan genes and the incompatibility of bacterial, archaean, and eukaryotic genetic systems, the tree model of evolution goes extinct. Evolutionary biologists, who waste most of their time contemplating line diagrams connecting various creatures, will then be free to go back to school to learn a useful trade.
Animal and plant classification can return to describing what living things are; each one unique.
The rest of the story
|
The late Stephen Jay Gould was one of the most influential evolutionary biologists of the 20th century and perhaps the best known since Charles Darwin, according to his New York Times obituary. In 1996 he wrote an essay about a famous giraffe evolution story in his "Natural History" magazine column. |
"I made a survey of all major high-school textbooks in biology. Every single one - no exceptions - began its chapter on evolution by first discussing Lamarck's theory of the inheritance of acquired characters, and then presented Darwins theory of natural selection as a preferable alternative. All texts then use the same example to illustrate Darwinian superiority - the giraffe's neck. Giraffes, we are told, got long necks in order to browse the leaves at the tops of acacia trees... available to no other mammal." "Darwinian evolution may be both true and powerful, but if we continue to illustrate our conviction with an indefensible, unsupported, entirely speculative, and basically rather silly story, then we are clothing a thing of beauty in rags - and we should be ashamed". "If we choose a weak and foolish speculation as a primary textbook illustration then we are in for trouble".
Although acacia tree leaves are the preferred food for adult giraffes during the wet season, giraffes will browse on many other trees and bush types. There is plenty of foliage at lower-levels, and giraffes often eat bushes and even low-growing land vegetation. They commonly munch on long grass and low bushes and many kinds of ground-growing plants.
The neck of the average female giraffe is two feet shorter than male necks. If, during a drought, only a longer neck could reach the last leaves high up on acacia trees, then the females would have starved to death and giraffes would have gone extinct.
Gould continues: "Even if we assume that the giraffe's neck evolved as an adaptation for eating high leaves, how could natural selection build such a structure by gradual increments? After all, the long neck must be associated with modifications in nearly every part of the body - long legs to accentuate the effect and a variety of supporting structures (bones, muscles, and ligaments) to hold up the neck. How could natural selection simultaneously alter necks, legs, joints, muscles, and blood flows (think of the pressure needed to pump blood to the giraffe's brain)?"
To drive blood eight feet up to the head, the heart is exceptionally large and thick-muscled, and the blood pressure is probably the highest in any animal. But when the giraffe bends its head to the ground it puts great strain on the blood vessels of the neck and head. The blood pressure plus the weight of the blood in the neck could produce so much pressure in the head that the blood vessels would burst. Pressure sensors along the neck's arteries monitor the blood pressure, and can activate other mechanisms to counter the increase in pressure as the giraffe drinks or grazes. Contracting artery walls (with increasing muscle fiber toward the head), shunting part of the blood flow to bypass the brain, and a web of small blood vessels (the rete mirabile, or "marvelous net") between the arteries and the brain all serve to control the blood pressure in the giraffe's head.
The lungs are oversize to compensate for the volume of dead air in the long trachea. Without this extra air-pumping capacity a giraffe would breathe the same used air over and over. The giraffe's lungs are very large and it breathes slowly, which is necessary in order to exchange the required large volume of air without causing windburn to the giraffe's 12 feet of trachea.
Red blood cells in a giraffe are about one-third the size of human red blood cells, so many more can fit into the same space. That provides giraffes with 3 times more red blood cell surface area than humans for the same volume of blood, producing higher and faster absorption of oxygen. This helps to retain adequate oxygen in all extremities, including the head.
Gould notes that "Giraffes provide no established evidence whatsoever for the mode of evolution of their undeniably useful necks." "Giraffes have a sparse fossil record in Europe and Asia and the spotty evidence gives no insight into how the long-necked modern species arose."
"The standard story, in fact, is both fatuous
and unsupported. In the realm of
giraffes, current use of maximal mamalian height for browsing leaves does not
prove that the neck evolved for such a function." "Why
then have we been bamboozled into accepting the usual tale without questioning? I suspect two primary reasons: we love a
sensible and satisfying story, and we are disinclined to challenge apparent
authority (such as textbooks)." --Gould,
Stephen Jay. May 1996. The Tallest Tale. Natural History, Vol. 105, Issue 5,
pp. 18-23, 54-57.
Giraffe biological
information from: Davis, Percival, and Dean H. Kenyon. 1993. Of Pandas and
People. Second edition, Haughton Publishing, Dallas, Texas.
An evolutionary science report takes down a creationist icon!
"If you want to see one of the wonders of the natural world, just startle a bombardier beetle. But be careful: when the beetles are scared, they flood an internal chamber with a complex cocktail of aromatic chemicals, triggering a cascade of chemical reaction that detonates the fluid and sends it shooting out of the insect's spray nozzle in a machine-gun-like pulse of toxic, scalding-hot vapor. The explosive, high-pressure burst of noxious chemicals doesn't harm the beetle, but it stains and irritates human skin - and can kill smaller enemies outright."
"The beetle's extraordinary arsenal has been held up by some as a proof of God's existence: how on earth, creationists argue, could such a complex, multistep defense mechanism evolve by chance? Now researchers at Stevens Institute of Technology in Hoboken, N.J. show how the bombardier beetle concocts its deadly explosives and in the process, learn how evolution gave rise to the beetle's remarkable firepower."
"Athula Attygalle, a research professor of chemistry and lead author of the work, said 'It turns out that the beetles' biochemistry is even more intricate than we'd thought.' Previously, researchers had assumed that two toxic, benzene-like chemicals called benzoquinones found in the beetles' spray were metabolized from hydroquinone, a toxic chemical that in humans can cause cancer or genetic damage. The team at Stevens showed that in fact just one of the beetle's benzoquinones derived from hydroquinone, with the other springing from a completely separate precursor: m-cresol, a toxin found in coal tar."
" 'It's fascinating that the beetles can safely metabolize such toxic chemicals', Attygalle said. In future studies, he hopes to follow the beetles' chemical supply chain further upstream, to learn how the precursors are biosynthesized from naturally available substances."
"The team's findings also show that the beetles' explosives rely on chemical pathways found in many other creepy-crawlies. Other animals such as millipedes also use benzoquinones to discourage predators, although they lack the bombardier's ability to detonate their chemical defenses. Evolutionarily distant creatures such as spiders and millipedes use similar strategies, too, suggesting that multiple organisms have independently evolved ways to biosynthesize the chemicals."
" 'That's a reminder that the bombardier beetle, though remarkable, is part of a rich and completely natural evolutionary tapestry', Attygalle said. 'By studying the similarities and differences between beetles' chemistry, we can see more clearly how they and other species fit together into the evolutionary tree', he explained. 'Beetles are incredibly diverse, and they all have amazing chemical stories to tell.' " - Benios, Thania. June 16, 2020. Research reveals the chemistry behind the bombardier beetle's extraordinary firepower. https://phys.org/news/2020-06-reveals-chemistry-bombardier-beetle-extraordinary.html
Well, did you learn how evolution gave rise to the beetle's remarkable firepower? No? Here is a hint: other unrelated insects use the toxic chemicals too; the bombardier beetle is not the only one. So we can say it evolved! If it were the only one, it would have evolved too, but this is better because all of them evolved. And that's how evolution did it by evolving. What? You want step by step genetic changes leading to complex novel function? Sorry; this is all you get from evolutionary science.
As an evolutionary biologist wrote, "the processes underlying evolutionary innovation are remarkably poorly understood, which leaves us at a surprising conundrum: while biologists have made great progress over the past century and a half in understanding how existing traits diversify, we have made relatively little progress in understanding how novel traits come into being in the first place." "The origin of novel features continues to be a fascinating and challenging topic in evolutionary biology."-- Moczek, Armin P. May 2008. On the origins of novelty in development and evolution. BioEssays, Vol. 30, Issue 5, pp. 409-512.
Evolve this: Blue morpho butterfly





Before the scientific era, people often made up imaginative stories to explain what they saw in the world. The scientific method changed that by requiring rigorous experimentation to test hypotheses and determine what is real. With the Theory of Evolution, people are back to making up imaginative stories. These excerpts from How Did Insect Metamorphosis Evolve? in Scientific American, August 10, 2012 by Ferris Jabr are a great example:
"Insects may account for between 80 and 90 percent of all animal species, which means 45 to 60 percent of all animal species on the planet are insects that undergo complete metamorphosis according to one estimate."
"However metamorphosis evolved, the enormous numbers of metamorphosing insects on the planet speak for its success as a reproductive strategy. The primary advantage of complete metamorphosis is eliminating competition between the young and old. Larval insects and adult insects occupy very different ecological niches. Whereas caterpillars are busy gorging themselves on leaves, completely disinterested in reproduction, butterflies are flitting from flower to flower in search of nectar and mates. Because larvas and adults do not compete with one another for space or resources, more of each can coexist relative to species in which the young and old live in the same places and eat the same things. Ultimately, the impetus for many of life's astounding transformations also explains insect metamorphosis: survival."
In fossils found in Permian rock, "some insects hatched in forms that neither looked nor behaved like their adult versions." This "incomplete metamorphosis, describes insects such as cockroaches, grasshoppers and dragonflies that hatch as nymphs--miniature versions of their adult forms that gradually develop wings and functional genitals as they molt and grow." " insects that mature through incomplete metamorphosis pass through a brief stage of life before becoming nymphs--the pro-nymphal stage, in which insects look and behave differently from their true nymphal forms."
 And
now, storytime:
And
now, storytime:
" the evolution of insect metamorphosis remains a genuine biological mystery even today." "Metamorphosis is a truly bizarre process". Nevertheless, "biologists have established a plausible narrative about the origin of insect metamorphosis, which they continue to revise as new information surfaces."
"Complete metamorphosis likely evolved out of incomplete metamorphosis." It "likely involved a genetic tweak that bathed the embryo in juvenile hormone sooner than usual and kept levels of the hormone high for an unusually long time."
"Perhaps 280 million years ago, through a chance mutation, some pro-nymphs failed to absorb all the yolk in their eggs, leaving a precious resource unused. In response to this unfavorable situation, some pro-nymphs gained a new talent: the ability to actively feed, to slurp up the extra yolk, while still inside the egg. If such pro-nymphs emerged from their eggs before they reached the nymphal stage, they would have been able to continue feeding themselves in the outside world. Over the generations, these infant insects may have remained in a protracted pro-nymphal stage for longer and longer periods of time, growing wormier all the while and specializing in diets that differed from those of their adult selves--consuming fruits and leaves, rather than nectar or other smaller insects. Eventually these prepubescent pro-nymphs became full-fledged larvae that resembled modern caterpillars." "The pupal stage arose later as a kind of condensed nymphal phase that catapulted the wriggly larvae into their sexually active winged adult forms."
|
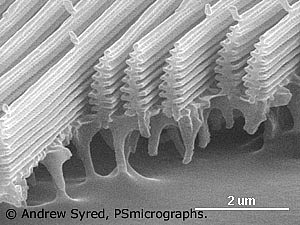
But wait - theres more! The underside of the wing has a brown pigment, which helps hide the resting blue morpho. |
|
|
|
That shimmering blue on top is not pigment. These extremely tiny shapes that cover the scales on top of the wing cause light wave interference. Blue light has a wavelength range from 400 to 480 nm. The slits in the scales of the Morpho are 200 nm apart. Because the distance between slits corresponds to half of the wavelength of blue light, this is the wavelength that undergoes constructive interference. The slits are attached to a base of melanin, a material that absorbs light, further strengthening the blue image. If evolutionists get around to making up a story for how these structures evolved, what do you think it will be? Come on, use your imagination! |
|
|
...or this: Pufferfish nests
|
|
The pufferfish in the video did not learn how to do this, it is hardwired in his brain. Can you guess which mutations occurred to build this unique behavior into the mind of a pufferfish? If you can, be sure to tell an evolutionary biologist; they need your help. |
Small pufferfish make a particular design in the sand off the coast of the Ryukyu Islands.
This species of pufferfish is less than 5 inches long, yet the male makes a circular structure 5 to 7 feet in diameter in seafloor sand over 7 to 9 days.
A female releases her eggs into the central zone. After spawning, males remain in the circular structure for 6 days to care for the eggs. Once the eggs hatch, males leave, never to return. But they begin to construct a new circular structure in a different place.
"The nest exhibits 3 unusual characteristics that have never been reported in fish. First, radially aligned peaks and valleys are created outside the nest site; second, the peaks are decorated with shell fragments; and third, fine sand particles are gathered in the nest site to create an irregular pattern. All 3 characteristics are completed and maintained before mating, when females visit the nest site, and they collapse thereafter."--Kawase, Hiroshi, Yoji Okata, Kimiaki Ito. 1 July 2013. Role of Huge Geometric Circular Structures in the Reproduction of a Marine Pufferfish. Scientific Reports, Vol. 3, Article number: 2106. 5 pages. DOI:10.1038/srep02106.
Can you begin to imagine the coded information in a pufferfish's DNA that produces this behavior? Evolution cannot write coded information.
...or this: Cuttlefish skin
|
|
|
Cuttlefish have "one of the most complex systems of motor coordination ever recorded." "Cuttlefish skin contains millions of cells called chromatophores, which can produce tiny dots of colour (yellow, orange, red, brown or black). If the radial muscles that control a chromatophore are relaxed, the pigments are imperceptible. But muscle contraction produces a colorful pixel several tens of micrometres wide." "The millions of individual pixels form a complex image". |
|
|
|
Cuttlefish transfix their prey by strobing as they approach. "Chromatophores are regulated by modules of motor neurons that function in synchrony, and that operate on skin patches of different sizes." There is "a remarkable level of fine control by motor neurons, and highlights the potential of cuttlefish studies to deepen our understanding of complex motor systems." |
The difference in
colour reflects a difference in age. The pigment of every chromatophore starts
as yellow before turning red, then brown, and ending up as black. New chromatophores
are generated throughout the life of the cuttlefish, and
the ratio of black to
colored chromatophores is maintained by keeping a tight balance between the
birth rate of new cells and the time it takes them to mature to a black color."
"The next challenge will be to determine how
cuttlefish change the 3D texture of their skin for camouflage on sand, algae or
corals. This process involves sets of muscles called papillae that create bumps
and lumps." "Cuttlefish coordinate millions
of muscles simultaneously".
Jouary, Adrien,
Christian K. Machens. 18 October 2018. A living display system.
Nature, Vol. 562, pp. 350-351.
Can you begin to imagine the coded information in a cuttlefish's DNA that produces this capability and behavior? Evolution cannot write coded information.
...or this: Scallop eyes (Quotes from a 2017 report in the journal Science)
|
|
|
Scallops possess a visual system comprising up to 200 eyes. What benefit does the scallop receive by having up to 200 eyes located on the periphery of its semi-circular mantle, spanning ~250°? The optic nerves from nearly all of the eyes project on to the site of visual processing in scallops. There, the scallop can combine the visual information from the... overlapping and differently focused views from multiple eyes. |

Each eye is ~1 mm in diameter and is composed of a cornea, a weakly refracting lens, and a concave mirror, in addition to a highly unusual double-layered retina.

Two striking features were observed in all the eyes. First, the mirror does not have a simple hemispherical shape. Rather, the curvature of the mirror varies across its surface. Second, the optical axes of the mirror and the lens are not aligned.
The mirror is tuned to reflect the wavelengths of light penetrating the scallops habitat and is tiled with a mosaic of square guanine crystals.
The crystals are arranged so that the high-refractive-index faces are oriented toward the direction of the incident light across the mirror, creating a highly reflective surface. The square-plate morphology is also optimized for tiling. Each layer of the mirror is formed from an almost perfectly tessellated mosaic of two-dimensional (2D) squares - closely resembling the segmented mirrors used in reflecting telescopes.
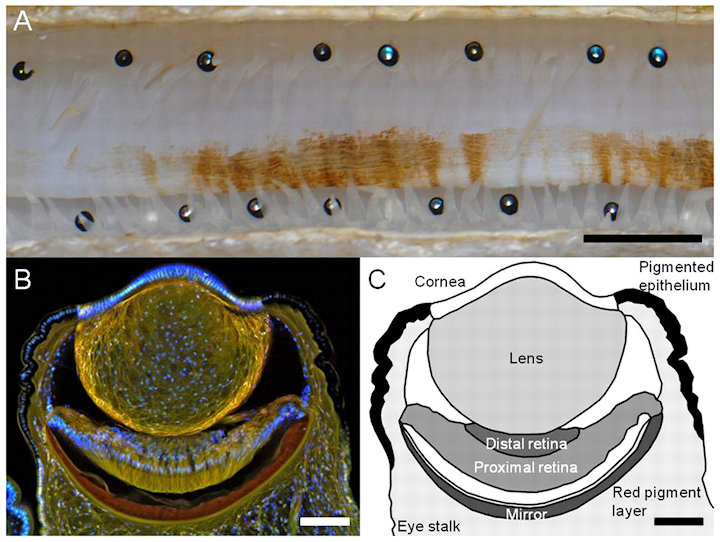
The multilayered mirror is constructed from 20 to 30 layers of crystals separated by thin layers of cytoplasm. Crystal tiling minimizes surface defects at the crystal interfaces that would cause optical diffraction effects.
The mirror forms images on a double-layered retina used for separately imaging the peripheral and central fields of view. The mirror forms functional images on both retinas, which appear to be specialized for different functions.
The distal retina responds to relatively dark, moving features, triggering defense or escape reflexes. The scallops well-focused peripheral vision could provide useful information to control and guide its movement.
Palmer, Benjamin A., Gavin J. Taylor, Vlad Brumfeld, Dvir Gur, Michal Shemesh, Nadav Elad, Aya Osherov, Dan Oron, Steve Weiner, Lia Addadi. 1 December 2017. The image-forming mirror in the eye of the scallop. Science Vol. 358 Issue 6367, pp. 1172-1175. DOI: 10.1126/science.aam9506
Can you begin to imagine the coded information in a scallop's DNA that produces these eyes? Evolution cannot write coded information.
or this: Orb weaver spider
|
|
|
People can choose many ways of life, inventing appropriate tools, machines, and techniques useful to survive and thrive. But other creatures are locked into their lifestyles, and are fully equipped for them. Evolutionists say living things "adopted strategies" as if they had a choice. Animals "adopting strategies" is as impossible today as it was in the deep mists of time. Design is the only objective conclusion. |
Look at the orb weaver spider. It is not taught or shown how to build a web; the skill, knowledge, and understanding is in its DNA. It knows how to choose a location, find anchor points in a plane for anchor threads, bind the anchor threads with frame threads, and run radius threads to a center point. It knows how to use its biological machines to make 4 to 7 different types of silk, including sticky silk for the capture spiral.
It knows to leave an open area in the center and to wait there. It knows to respond to vibrations of the radius threads when an insect is caught. It knows to walk only on the radius threads so it won't get caught on the sticky capture threads. It knows how to wrap and devour a captured insect, and to repair the web.
Robotics engineers struggle to design robots that can keep their balance or grasp a cup without crushing it. The extremely complex, precise, and purposeful behavior of orb weaver spiders is light-years beyond robotics programming, and it's all due to coded information in spider DNA. Evolution cannot write coded information.
...or this: Lampsilis mussel fooling bass
|
|
The Lampsilis mussel cannot see.
The lure is startling, but also remember that the mussels microscopic larvae (glochidia) must become parasites in bass for a while in order to develop. Can you begin to imagine the coded information in Lampsilis mussel DNA that produces this larva and lure? Evolution cannot write coded information. Image from: https://www.youtube.com/watch?v=I0YTBj0WHkU |
...or this: a daisy fooling flies
|
|
"An orange-hued daisy in South Africa has an unusual lure to attract pollinators: a little structure on its petals that resembles a female fly. Male flies descend on the petals in hopes of mating but end up ferrying the flowers' pollen to other plants." Wetzel,
Corryn. 23 March 2023. How daisies make
deceptive petals that look like female flies. NewScientist, Life. https://www.newscientist.com/article/2365794-how-daisies-make- |
|
|
The daisies are pollinated by the "bombyliidae fly (Megapalpus capensis Wiedeman)". "Particularly sophisticated complex traits are involved in plant sexual deception, when flowers evolve novel structures mimicking mating signals of female insects to attract males for pollination." "The evolution of sexually deceptive flowers thus necessitates orchestrated changes in several genetic networks altering multiple unrelated floral features". Kellenberger et al. 2023. Multiple gene co-options underlie the rapid evolution of sexually deceptive flowers in Gorteria diffusa. Current Biology, Vol. 33, pp. 1-11 DOI:10.1016/j.cub.2023.03.003 |
The evolutionist authors believe "that the rapid evolution of sexual deception in G. diffusa was propelled by independent co-options of genetic elements affecting the pigmentation, cellular structure, and spatial organization of pre-existing petal spots."
They imagine that there was "excessive tinkering with independently co-opted genetic modules". Wow! Who did that? Nobody! In the evolutionist mind, tinkering just happens; excessive tinkering.
For those of us in the real world, this is an obvious case of engineering using coded information that Evolution theory cannot produce.
or this: a worm controlling the mind of a praying mantis|
|
Horsehair (or gordian worms) are a group of parasitic animals.
Many have complex life cycles involving multiple hosts, and the
ones that live in freshwater must generally find their way into an
insect to finish developing into adults.
The genus Chordodes infect mantises and can grow to nearly 1 meter long inside their abdomens. Once they've finished growing, the worms must convince their hosts to drown themselves to complete the worms' life cycle. Researchers found more than 3100 parasite genes with increased expression while they were controlling their hosts. More than 1400 of these worm genes closely matched genes from the mantises. |
It's possible that the worms' proteins may mimic those of their hosts, allowing the worms to commandeer the mantids' bodies, activating behavioral programs that suit their needs.-- Wilcox, Christie. Parasitic worms may control minds of insects with 'borrowed' genes. Science 19 Oct 2023 11:45 AM ET News DOI: 10.1126/science.adl4678
Inside cells
|
|
An adult human is made up of approximately 100 trillion cells. Every cell contains approximately one billion protein molecules that perform different important functions. Even a simple yeast cell is made up of roughly 42 million proteins. The proteins within a cell are constantly degraded and resynthesized (replaced). |
In the human genome, only about 1% of our DNA is genes. Most genes contain the information needed to make proteins. A few genes produce regulatory molecules that help the cell assemble proteins. Our bodies make somewhere between 80,000 and 400,000 different types of proteins, each with a specific function.
The rest of our DNA governs every bodily activity.
|
|
Every cell contains many different organelles that are specialized to carry out different tasks, each surrounded by a membrane. A protein carries in its structure the information needed to specify its proper location in the cell. These signals can be compared to address tags or zip codes which ensure that a travelers luggage arrives at the correct destination, or a letter reaches its correct addressee. Signal sequences are either a short "tail" at one end of the protein or located within the protein. Specific amino acid sequences (topogenic signals) determine whether a protein will pass through a membrane into a particular organelle, become integrated into the membrane, or be exported out of the cell. It all operates the same way in yeast, plant, and animal cells. https://www.nobelprize.org/prizes/medicine/1999/press-release/ |
The number
of amino acids that make up all proteins range from as few as 50 up to several
thousand, forming long, folded chains. This video
summarizes how protein coding instructions are used in you and me.
From: https://www.youtube.com/watch?v=gG7uCskUOrA
and https://www.youtube.com/watch?v=vi-zWoobt_Q
|
Now blow your mind with a video of what is happening constantly in your cells at real-time speed, and imagine Darwin's reaction if he saw it. |
|
| |
The rough endoplasmic reticulum, studded with millions of membrane bound ribosomes, is involved with the production, folding, quality control and dispatch of some proteins. Some of the proteins are delivered into the lumen (space inside the endoplasmic reticulum) while others are processed within the endoplasmic reticulum membrane itself. It is in the lumen of the rough endoplasmic reticulum that proteins are folded to produce the highly important biochemical architecture which will provide "lock and key" and other recognition and linking sites. |
It is also in the lumen that an amazing process is carried out. Proteins are subjected to a quality control check, and any that are found to be incorrectly formed or incorrectly folded are rejected. These rejects are stored in the lumen or sent for recycling for eventual breakdown to amino acids.
| |
In most
cases proteins are transferred to the Golgi apparatus for "finishing". They are conveyed in vesicles or possibly
directly between the endoplasmic reticulum and Golgi surfaces. After
"finishing" they are delivered to specific locations. https://bscb.org/learning-resources/softcell-e-learning/endoplasmic- |
Natural selection could only work on a gene-copying system that already exists; it could not invent such a system. The fact that Evolution theory has no way to invent the protein-making system that is in all plants and animals, and no mind to invent coded information, proves that Evolution theory has no connection with reality.
The fuel for cells to keep working, ATP, is another awesome design:
Gradual change
versus leaps
There are two versions of evolution
theory. The
main version proposes that many tiny changes over millions of years made new
creatures. It is called the Modern Synthesis or Neo-Darwinian evolution.
But "major transitions in biological evolution show the same pattern of sudden emergence of diverse forms at a new level of complexity." "The principal 'types' seem to appear rapidly and fully equipped with the signature features of the respective new level of biological organization. No intermediate 'grades' or intermediate forms between different types are detectable."23
Since the fossil record does not show long series of tiny changes between one type of creature and another, some evolutionists proposed a modification to evolution theory. It says that change occurred by occasional leaps (punctuated equilibrium), not gradually. However, each hypothetical beneficial mutation could only make a slight change. Any more than that would be so disruptive as to cause death. So punctuated equilibrium is not really about big leaps. It envisions a lot of slight changes over thousands of years, then nothing happens for millions of years. Evolutionists say with a straight face that no fossils have been found from a leap because thousands of years is too fast in the billions of years of "geologic time" to leave any. Yet without fossils there is no evidence that any leaps ever happened, and of course there is no evidence that leaps or gradual changes beyond variation are happening today in any of the millions of species that still exist.
Evolutions Third Way
"Evolution
is changed gene frequencies in populations." - Richard Dawkins
Evolution theory says that accumulated small changes in creatures (microevolution) lead to new types of creatures (macroevolution). But some evolutionary biologists are admitting that microevolution does not happen by the supposed mechanism of evolution - mutation/natural selection. Instead, living things have built-in mechanisms that adjust to quick changes in their environment to produce variation. The mechanisms are only beginning to be understood, yet 77 evolutionist academics have put their names and faces on The Third Way website.
A system for variation makes sense because species' survival can depend on adapting fast and not waiting millions of years for "beneficial mutations". But this leaves macroevolution out hanging by itself, which is why Third Way members are often bitterly opposed by conventional Neo-Darwinists. The quotes below were on The Third Way website; they have since been removed: http://www.thethirdwayofevolution.com/
|
"New findings in molecular biology challenge the gene-centered version of Darwinian theory according to which adaptation occurs only through natural selection of chance DNA variations." "The DNA record does not support the assertion that small random mutations are the main source of new and useful variations. We now know that the many different processes of variation involve well regulated cell action on DNA molecules." " the twentieth-century scientific consensus about evolution appears outdated and incomplete due to the inadequacy of natural selection and adaptation as the only or even the main mode of evolution". "The fossil record, in fact, does not show Darwin's predicted gradual changes between closely related species but rather the "punctuated equilibrium" pattern described by Eldredge and Gould: a jump from one to a different species." "How do new species evolve? Although Darwin identified inherited variation as the creative force in evolution, he never formally speculated where it comes from. His successors thought that new species arise from the gradual accumulation of random mutations of DNA. But despite its acceptance in every major textbook, there is no documented instance of it." "The genes eye view of life, advocated by evolutionary biology, sees living bodies as mere vehicles for the replication of the genetic codes." But "understanding the components of a system (be they individual genes, proteins, or even molecules) may tell us little about the interactions among these components." "Neo-Darwinism ignores much contemporary molecular evidence and invokes a set of unsupported assumptions about the accidental nature of hereditary variation. Neo-Darwinism ignores important rapid evolutionary processes such as symbiogenesis, horizontal DNA transfer, action of mobile DNA and epigenetic modifications. Moreover, some Neo-Darwinists have elevated Natural Selection into a unique creative force that solves all the difficult evolutionary problems without a real empirical basis." "Evolution, as it turns out, is much more dynamic than biologists realized just a few decades ago. Genomes merge, shrink and grow, acquire new DNA components, and modify their structures by well-documented cellular and biochemical processes." " evolutionary change [is] an active cell process, regulated epigenetically and capable of making rapid large changes by horizontal DNA transfer, inter-specific hybridization, whole genome doubling, symbiogenesis, or massive genome restructuring." "To understand what life is, we must view it at a variety of different levels, all interacting with each other in a complex web. It is that emergent web, full of feedback between levels, from the gene to the wider environment, that is life." |
Fossil record
https://www.youtube.com/watch?v=20AGi50UNf0
Fossils are what is left of living things that were buried quickly. They range from impressions to mineral
replacements to decayed organics.
Evolutionary biologists describe differences between living things and
then make up stories about them. In 2022
a revered story about bird beaks got overturned:
"Each of the roughly 11,000 species of birds on Earth today is classified into one of two over-arching groups, based on the arrangement of their palate bones. Ostriches, emus and their relatives are classified into the palaeognath, or 'ancient jaw' group, meaning that, like humans, their palate bones are fused together into a solid mass."
"All other groups of birds are classified into the neognath, or 'modern jaw' group, meaning that their palate bones are connected by a mobile joint. This makes their beaks much more dexterous, helpful for nest-building, grooming, food-gathering, and defense."
"The two groups were originally classified by Thomas Huxley, the British biologist known as 'Darwins Bulldog' for his vocal support of Charles Darwin's theory of evolution. In 1867, he divided all living birds into either the 'ancient' or 'modern' jaw groups. Huxleys assumption was that the 'ancient' jaw configuration was the original condition for modern birds, with the 'modern' jaw arising later."
"'This assumption has been taken as a given ever since,' said Dr. Daniel Field from Cambridge's Department of Earth Sciences". That is, until now.
|
|
This is Janavis finalidens, discovered in late Cretaceous strata. "Researchers from the University of Cambridge and the Natuurhistorisch Museum Maastricht found that one of the key skull features that characterizes 99% of modern birds a mobile beak evolved before the mass extinction event that killed all large dinosaurs, 66 million years ago." |
"'Evolution doesn't happen in a straight line,' said Field. 'This fossil shows that the mobile beak a condition we had always thought post-dated the origin of modern birds, actually evolved before modern birds existed. We've been completely backwards in our assumptions of how the modern bird skull evolved for well over a century.'"-- Collins, Sarah. 30 November 2022. Fossil overturns more than a century of knowledge about the origin of modern birds. University of Cambridge news online: https://www.cam.ac.uk/stories/the-last-toothed-bird
No big deal; the assumptions of Evolution theory lead to false conclusions, so evolutionary biologists change their stories from time to time and call it progress. Since they can't begin to fathom how gene regulatory networks could be altered or how new proteins could be devised to accomplish the changes they claim occurred through random mutation, they just say the magic words "it evolved" and everybody nods their heads knowingly.
Evolution is supposed to be all about change, whether gradual or in leaps. Consider a cloud in the sky: it is constantly changing shape due to natural forces. Perhaps it looks like a rabbit now, and a few minutes later like a horse. In between, the whole mass is shifting about. In a few more minutes it may look like a bird.
The problem for evolution is that we never see the shifting between shapes in the fossil record. All fossils are of complete animals and plants, not works in progress "under construction". That is why we can give each distinct plant or animal a name.
If evolution's continuous morphing were really going on, every fossil would show change underway throughout the creature, with parts in various stages of completion. For every successful change there should be many more that lead to nothing. The whole process is random trial and error, without direction. So every plant and animal, living or fossil, should be covered inside and out with useless growths and have parts under construction. It is a grotesque image, and just what the theory of evolution really predicts.
Even Charles Darwin had a glimpse of the problem in his day. He wrote in his book On the Origin of Species: "The number of intermediate varieties which have formerly existed on Earth must be truly enormous. Why then is not every geological formation and every stratum full of such intermediate links? Geology assuredly does not reveal any such finely graduated organic chain; and this, perhaps, is the most obvious and gravest objection which can be urged against my theory."
The more fossils that are found, the better sense we have of what lived in the past. Since Darwin's day, the number of fossils that have been collected has grown tremendously, so we now have a pretty accurate picture. The gradual morphing of one type of creature to another that evolution predicts is nowhere to be found. There should have been millions of transitional creatures if evolution were true. In the "tree of life" that evolutionists have dreamed up, gaps in the fossil record are especially huge between single-cell creatures, complex invertebrates (such as snails, jellyfish, trilobites, clams, and sponges), and what evolutionists claim were the first vertebrates, fish.
In fact, there are no transitional fossils at all between single-celled creatures and complex invertebrates, nor between complex invertebrates and fish. That alone is fatal to the theory of evolution. The fossil record shows that evolution never happened.
What fossil evidence is there for the evolutionist vision for the origin of life? Nothing, except for tiny filaments, knobs and tubes in Canadian rocks supposedly 4.28 billion years old.
Everyone agrees that the big surprise is the sudden appearance of fossils above the bedrock in the Cambrian Explosion, supposedly 541 to 530 million years ago.
|
|
|
The fossils of the Cambrian Explosion are complex invertebrates, sea creatures like trilobites, sponges, worms, jellyfish, sea urchins, sea lilies, mollusks, brachiopods (lamp shells), sea cucumbers, and swimming crustaceans such as |
|
|
|
Opabinia, 3 inches long (8 cm) with 5 eyes and a long claw arm, and |
|
|
|
Anomalocaris, 3 feet long (91.5 cm), and the top predator in the Cambrian environment. |
"Darwin argued that the incompleteness of the fossil record gives the illusion of an explosive event, but with the eventual discovery of older and better-preserved rocks, the ancestors of these Cambrian taxa would be found. Studies of Ediacaran and Cambrian fossils continue to expand the morphologic variety of clades, but the appearance of the remains and traces of bilaterian animals in the Cambrian remains abrupt."-- Erwin, Douglas H., Marc Laflamme, Sarah M. Tweedt, Erik A. Sperling, Davide Pisani, Kevin J. Peterson. 2011. The Cambrian Conundrum: Early Divergence and Later Ecological Success in the Early History of Animals. Science, Vol. 334, pp. 1091-1097.
Abrupt indeed. Here is "the
ancestor of all animals" supposedly from 14 million years
before even the Cambrian Explosion: "Geologists have
discovered the first ancestor on the family tree that contains most
animals today, including humans. The worm-like creature, Ikaria
wariootia, is the earliest bilaterian, or organism with a front
and back, two symmetrical sides, and openings at either end connected
by a gut. It was found in Ediacaran Period deposits in Australia
and was 2 to 7 millimeters long, with the largest the size of a
grain of rice."
-- https://www.sciencedaily.com/releases/2020/03/200323152108.htm
|
|
|
In the classification diagram biologists use, all these animals are so unrelated to each other that they are in different classes or even phyla. From time to time evolutionists announce with great fanfare that they have gotten a colony of bacteria to eat something they could not eat before, or some other small variation. These changes are always below the family level on the diagram. If evolution were true, there would have been ancestors and transitional creatures between each genus, family, order, class, and phylum in the layers below the Cambrian Explosion. But there are no fossils for any of these. |
What to do? A team of evolutionists solved this problem using their most effective tool - storytelling.
First they assumed evolution occurred. Then they estimated how fast it should have happened, and decided that the creatures in the Cambrian Explosion had been evolving for over 250 million years before any showed up in the rocks as fossils!
"We estimate that the last common ancestor of all living animals arose nearly 800 million years ago and that the stem lineages leading to most extant phyla had evolved by the end of the Ediacaran (541 million years ago)."
Yes, millions of generations of all kinds of creatures all over the world living, dying, evolving, without leaving any trace of their existence.
Not only that, "from the early Paleozoic onward there is little addition of new phyla and classes". "Little high-level morphological innovation occurred during the subsequent 500 million years". Their story was published in the prestigious journal Science, and hailed as having solved a mystery challenging evolution theory all the way back to Darwin.-- Erwin, Douglas H., Marc Laflamme, Sarah M. Tweedt, Erik A. Sperling, Davide Pisani, Kevin J. Peterson. 2011. The Cambrian Conundrum: Early Divergence and Later Ecological Success in the Early History of Animals. Science, Vol. 334, pp. 1091-1097.
That conclusion was based on principles of Evolutionary Science, so it is no surprise that hard evidence proves it wrong.
"The apparent clash between molecular clock estimates of the time of origins of clades versus what the fossil record might be thought to say has been a significant source of anxiety in the last two decades."
"Bayesian molecular clock analyses are popular because they can be run on large datasets and can take into account many uncertainties by setting priors over them. However, the calibrations, and, in particular, the priors that arise from them, are typically not biologically informed."
"Molecular clock estimates, which have sometimes been argued to have essentially taken over from the fossil record as the ultimate measures of evolutionary time, conversely, often give much larger gaps between first appearances of clades and the fossil record."
"The ingrained idea that the fossil record can only start after a long and unpredictable lag period or phylogenetic fuse has been criticized on various grounds, but seems to be the basis for the relative insouciance [unconcern] with which large gaps between clocks and fossils are accepted."
" the molecular part of the analysis does not allow us to distinguish between different times of origin of the clade. the fossil record can provide a powerful test of molecular clock methodology, and why it goes astray, and we have every reason to think these problems are general."-- Budd, Graham E., Richard P. Mann. 11 September 2023. Two Notorious Nodes: A Critical Examination of Relaxed Molecular Clock Age Estimates of the Bilaterian Animals and Placental Mammals. Systematic Biology, syad057, DOI:10.1093/sysbio/syad057
Trilobites are at the bottom of the fossil record, along with other creatures in the Cambrian explosion.
|
|
"Darwin chose trilobites as an exemplar group to highlight his dilemma about animal origins:" |
|
|
|
"There is another and allied difficulty, which is much graver. I allude to the manner in which numbers of species of the same group, suddenly appear in the lowest known fossiliferous rocks... For instance, I cannot doubt that all the [Cambrian] trilobites have descended from some one crustacean, which must have lived long before the [Cambrian] age (p. 306)." |
"Here we test Darwins hypothesis", and the "dataset is the largest and most comprehensive for trilobites compiled to date".
"We conclude that the Cambrian explosion was over by the time the typical Cambrian fossil record commences and reject an unfossilized Precambrian history for trilobites".
Again: "Our data therefore provide robust, quantitative
evidence that by the time the typical Cambrian fossil record begins (~521 Ma), the Cambrian
explosion had already largely concluded."
-- Paterson, John R., Gregory D.
Edgecombe, and Michael S. Y. Lee. March 5, 2019. Trilobite evolutionary rates
constrain the duration of the Cambrian explosion. Proceedings of the National
Academy of Sciences (PNAS), Vol. 116, No. 10, pp. 43944399. DOI: 10.1073/pnas.1819366116
But wait; theres more!
"Among the various trilobite groups, one trilobite, Dalmanitina socialis, possessed a unique visual system with compound eyes composed of two optically homogeneous lens units of different refractive indices an upper lens unit with a central bulge made of calcite and a lower lens unit made of an organic compound. As a result, each compound eye of Dalmanitina socialis is able to simultaneously focus incident light to a near and a far point", and simultaneously perceive both close and distant objects".
" this type of compound-eye visual system is unique to Dalmanitina socialis, and is in contrast to the single focal vision system present in all-known living arthropods that exist today."
"Inspired by compound eyes of the trilobite Dalmanitina socialis, we design and
construct a chiral light-field camera incorporating an array of photonic
spin-multiplexed bifocal metalenses. Combined with a deep-learning-based neural
network reconstruction algorithm, the system provides distinct aberration-free
photographic capabilities, including the ability to achieve a polarization-controllable
extreme depth of field imaging while maintaining high spatial lateral
resolution."
Fan, Qingbin, Weizhu
Xu, Xuemei Hu, Wenqi Zhu, Tao Yue, Cheng Zhang, Feng Yan, Lu Chen, Henri J.
Lezec, Yanqing Lu, Amit Agrawal, Ting Xu. 19 April 2022. Trilobite-inspired neural nanophotonic
light-field camera with extreme depth-of-field. Nature Communications, Vol. 13,
No. 2130, 10 pages. DOI: 10.1038/s41467-022-29568-y
|
|
|
Fossil compound eyes from the Lower Cambrian, where the first complex creatures suddenly appear in the fossil record, have been found in the Emu Bay Shale of South Australia. The fossils are supposedly about 515 million years old. They may be corneas of Anomalocaris that were shed during moulting. The lenses are packed tighter than Lower Cambrian trilobite eyes, "which are often assumed to be the most powerful visual organs of their time." Notice that the lenses in the picture are different sizes. It is the same in the fossils. Each eye has "over 3,000 large ommatidial lenses". "The arrangement and size-gradient of lenses creates a distinct [forward] 'bright zone'... where the visual field is sampled with higher light sensitivity (due to larger ommatidia) and possibly higher accuity". This indicates "that these eyes belonged to an active predator that was capable of seeing in low light." "The eyes are more complex than those known from contemporaneous trilobites and are as advanced as those of many living forms" today, such as the fly in this picture, "revealing that some of the earliest arthropods possessed highly advanced compound eyes".27 When the earliest form is the most complex, there is no evolution. |
|
|
|
This tiny fish (a little over an inch long, or 3 cm) is Haikouichthys. Its fossils have also been found in the Lower Cambrian. This "first fish" has a spine and spinal cord, eyes, gills, fins, scales, mouth, etc., though no jaw, like a lamprey. About 500 were found buried together.39 |
|
|
|
This is Guiyu, a fossil fish that "represents the oldest near-complete gnathostome (jawed vertebrate)."48 It measures about 15 inches long, or 37 cm. Clearly, the earliest fish were as much fish as today's fish. Guiyu is "a representative of modern fishes" from the Silurian, before the so-called "age of fishes" (Devonian).9 In the evolutionist's mind, "a whole series of major branching events... must have taken place well before the end of the Silurian." "A significant part of early vertebrate evolution is unknown."9 |
|
|
|
Coelacanth disappeared from the fossil record with the last of the dinosaurs. That was supposedly 65 million years ago. In the early 1900s, evolutionists touted it as the first walking fish, the transition between fish and tetrapods. That is, until 1938 when one was found alive and unable to walk. Evolution theory says that pressures from competition and the environment force changes over time. In chapter 9 of his book, Darwin wrote of ancestor species in general: "If, moreover, they had been the progenitors of these orders, they would almost certainly have been long ago supplanted and exterminated by their numerous and improved descendants." Here is a coelacanth today, alive and unchanged like many "living fossils". Where is the evolution? |
|
|
|
Evolutionists tell us this dragonfly has not shown up in the fossil record for 250-300 million years! Dozens of the Ancient Greenling Damselfly live near Melbourne, Australia. "The damselfly, part of the dragonfly group Odonata, is the only living representative of the family Hemiphlebiidae. Its ancient predecessors are found solely in 250-300 million-year-old fossil records from Brazil to Russia." --Smith, Bridie. January 5, 2010. Found: fossil-linked, listed damselfly. www.theage.com.au (newspaper website) |
|
|
|
This is a drawing of a supposed predecessor, Protozygoptera. With a wingspan of under 6 cm, it is the earliest damselfly-like insect ever found and "the origin of modern dragonflies". Its fossil wing was found in rocks of the Upper Carboniferous which evolutionists think are about 300 million years old. As with many creatures, dragonflies appear suddenly in the fossil record, fully formed. Damselflies living today look like Protozygoptera; there are no transitional intermediates and there was no evolution. --Jarzembowski, E.A., A. Nel. 2002. The earliest damselfly-like insect and the origin of modern dragonflies (Insecta: Odonatoptera: Protozygoptera). Proceedings of the Geologists' Association, Vol. 113, pp. 165-169. |
|
|
|
Evolutionists always point to Archaeopteryx as the great example of a transitional creature, appearing to be part dinosaur and part bird. However, it is a fully formed, complete animal with no half-finished components or useless growths. Most people know "the stereotypical ideal of Archaeopteryx as a physiologically modern bird with a long tail and teeth". Research now "shows incontrovertibly that these animals were very primitive". "Archaeopteryx was simply a feathered and presumably volant [flying] dinosaur. Theories regarding the subsequent steps that led to the modern avian condition need to be reevaluated." --Erickson, Gregory, et al. October 2009. Was Dinosaurian Physiology Inherited by Birds? Reconciling Slow Growth in Archaeopteryx. PLoS ONE, Vol. 4, Issue 10, e7390. |
"Archaeopteryx has long been considered the iconic first bird." "The first Archaeopteryx skeleton was found in Germany about the same time Darwin's Origin of Species was published. This was a fortuituously-timed discovery: because the fossil combined bird-like (feathers and a wishbone) and reptilian (teeth, three fingers on hands, and a long bony tail) traits, it helped convince many about the veracity of evolutionary theory." "Ten skeletons and an isolated feather have been found." "Archaeopteryx is the poster child for evolution." But "bird features like feathers and wishbones have recently been found in many non-avian dinosaurs". "Microscopic imaging of bone structure... shows that this famously feathered fossil grew much slower than living birds and more like non-avian dinosaurs." "Living birds mature very quickly and grow really, really fast", researchers say. "Dinosaurs had a very different metabolism from today's birds. It would take years for individuals to mature, and we found evidence for this same pattern in Archaeopteryx and its closest relatives". "The team outlines a growth curve that indicates that Archaeopteryx reached adult size in about 970 days, that none of the known Archaeopteryx specimens are adults (confirming previous speculation), and that adult Archaeopteryx were probably the size of a raven, much larger than previously thought." "We now know that the transition into true birds -- physiologically and metabolically -- happened well after Archaeopteryx."--October 2009. Archaeopteryx Lacked Rapid Bone Growth, the Hallmark of Birds. American Museum of Natural History, funded science online news release.
What evolutionists now know for sure is that their celebrity superstar was not a transitional creature after all. Wow! OMG. They better find a new one fast...
|
|
|
How about the Platypus? They could call it a transitional creature between ducks and mammals. The furry platypus has a duck-like bill, swims with webbed feet, and lays eggs. |
As for the birds in the evolutionary tree, evolutionists just placed living and extinct species next to each other to make the bird series.
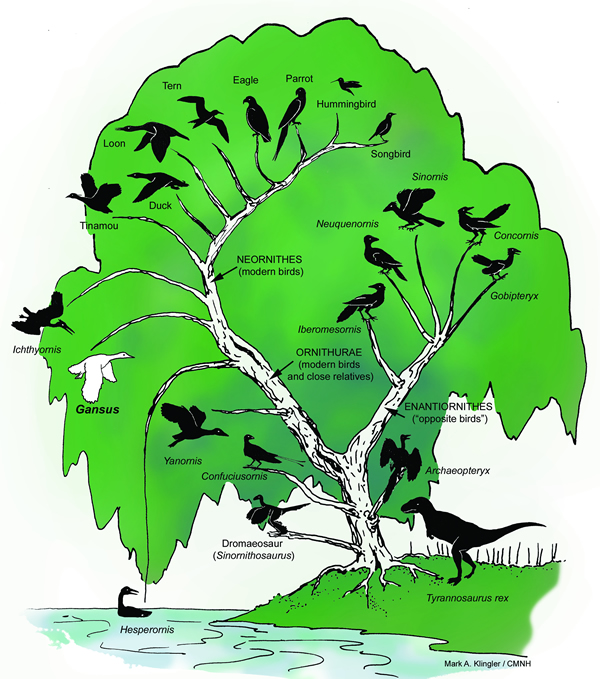
|
|
|
Early bird: Fossils of Archaeornithura meemannae were found in very Early Cretaceous strata in China. In the evolutionary tree (above) it sits at the bottom between Archaeopteryx and Confuciusornis. Do you see anything primitive here? "The fossils' specialized anatomy suggests that key factors in birds' long-term success, such as expert flying ability and rapid growth rates, arose surprisingly early in avian evolution" "and make it almost certain that the origin of the lineage was much older still."--Wang, Min, et al. 5 May 2015. The oldest record of ornithuromorpha from the early cretaceous of China. Nature Communications, 6:6987, DOI:10.1038/ncomms7987 |
|
|
|
The famous horse series; it looks great, doesn't it? But each of the supposed ancestors is a complete animal. They are not full of failed growths and there are no parts under construction. There are many more differences between each type of animal than their size and the number of toes. Every change in structure, function, and process would have had to develop through random trial-and-error if evolution were true, but no transitional forms have been found. The fossils have not caught any changes in the midst of being created, even though they should have occurred over long periods of time. In the late 1800's, evolutionists simply placed living and extinct species next to each other to make the horse series. However, evolutionists no longer believe there was the direct ancestry (orthogenesis) shown in this chart... |
|
|
|
Evolutionists now imagine it to be this branching bush. Many of the supposed ancestors apparently lived at the same time, especially after Mesohippus. It is doubtful that Hyracotherium (formerly Eohippus) has any connection to horses. So the progression of toes is an illusion that was useful when the theory of evolution was first being sold to the public. Several hundred species are extinct; only one genus, Equus, survives. |
Rather than play the evolutionist's game and try to untangle varieties of one animal from another in the horse bush, let's be clear on what we are talking about. Biologists divide all living things into groups and subgroups. The basic framework is the Linnaean system of taxonomy, published in Linnaeus' expanded 10th edition of Systema Naturae in 1758. That was a century before Darwinism, and it was never intended to show that one creature morphed into another. It just grouped animals with similar characteristics. Once they seized control of the study of biology, evolutionists took over the Linnaean system and have tinkered with it ever since to fit their belief that animals transform over time. Birds are at the class level (Aves), which has 23 subgroups below it called orders and 142 subgroups below them called families. All the members of the evolutionist's horse bush, living and extinct, are in one family, (Equidae). To get up to the class level where birds are, you pass the order Perissodactyla (browsing and grazing mammals with an odd number of toes) to the class mammals (Mammalia). Other examples of families include cats (Felidae), dogs (Canidae), deer (Cervidae), bears (Ursidae), squirrels (Sciuridae), and cattle (Bovidae).
So the level of "evolution" in the horse bush is within a family, which is pretty low and has no relevance to the main issue, macroevolution. But evolutionists have always used the varieties in the family Equidae to entice people to imagine that all animals morph from one kind to another. Look at their tree of life for mammals, the class Mammalia.
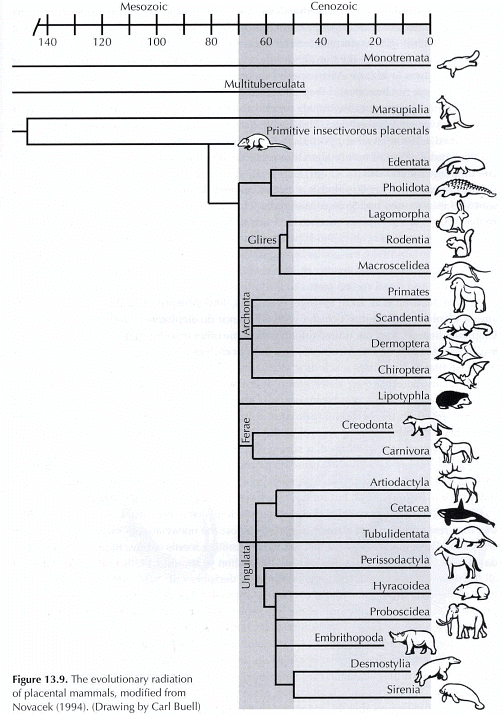
Novacek, M.J. 1994. The Radiation of Placental Mammals, in Prothero, D.R. and Schoch, R.M. (editors) Major Features of Vertebrate Evolution. Paleontological Society Short Courses in Paleontology, No. 7, pp. 220-237.
That's right; they want us to believe that elephants and manatees, primates and tree shrews had common ancestors. They show you three toes changing to one toe, browsing teeth changing to grazing teeth in the horse bush so that you will believe that reindeer and whales morphed from common ancestors. That is why the horse series is an evolutionist icon.
The "Tree of Life" is falling
|
|
Matthew Wills, Professor of Evolutionary Paleobiology at the Milner Center for Evolution at the University of Bath in England said: "It turns out that we've got lots of our evolutionary trees wrong. For over a hundred years, we've been classifying organisms according to how they look and are put together anatomically, but molecular data often tells us a rather different story." "Research led by scientists at the Milner Center for Evolution at the University of Bath suggests that determining evolutionary trees of organisms by comparing anatomy rather than gene sequences is misleading. The study, published in the journal Communications Biology on May 31, 2022, shows that we often need to overturn centuries of scholarly work that classified living things according to how they look." |
The study found that convergent evolution - when a characteristic evolves separately in two genetically unrelated groups of organisms - is much more common than biologists previously thought. So much for descent with modification...
Professor Wills said: "We already have lots of famous examples of convergent evolution, such as flight evolving separately in birds, bats, and insects, or complex camera eyes evolving separately in squid and humans. But now with molecular data, we can see that convergent evolution happens all the time - things we thought were closely related often turn out to be far apart on the tree of life.
| |
"It proves that evolution just keeps on re-inventing things, coming up with a similar solution each time the problem is encountered in a different branch of the evolutionary tree." "It means that convergent evolution has been fooling us - even the cleverest evolutionary biologists and anatomists - for over 100 years!" - Dr. Matthew Wills |
Ref: Oyston, Jack W., Mark Wilkinson, Marcello Ruta, Matthew A. Wills. Molecular phylogenies map to biogeography better than morphological ones. 31 May 2022. Communications Biology, Vol. 5, No. 521, pp. 1-12. DOI:10.1038/s42003-022-03482-x
Press release from: University Of Bath. Study suggests that most of our evolutionary trees could be wrong. June 1, 2022. https://www.bath.ac.uk/announcements/study-suggests-that-most-of-our-evolutionary-trees-could-be-wrong/
New discoveries are bringing down the whole notion of a "tree of life", as passages from an article in the mainstream magazine New Scientist show:26 "The tree-of-life concept was absolutely central to Darwin's thinking, equal in importance to natural selection, according to biologist W. Ford Doolittle of Dalhousie University in Halifax, Nova Scotia, Canada. Without it the theory of evolution would never have happened." "For much of the past 150 years, biology has largely concerned itself with filling in the details of the tree. 'For a long time the holy grail was to build a tree of life,' says Eric Bapteste, an evolutionary biologist at the Pierre and Marie Curie University in Paris, France. A few years ago it looked as though the grail was within reach."
"But today the project lies in tatters, torn to pieces by an onslaught of negative evidence. Many biologists now argue that the tree concept is obsolete and needs to be discarded. 'We have no evidence at all that the tree of life is a reality,' says Bapteste. That bombshell has even persuaded some that our fundamental view of biology needs to change." "The problems began in the early 1990s when it became possible to sequence actual bacterial and archaeal genes". "As more and more genes were sequenced, it became clear that the patterns of relatedness could only be explained if bacteria and archaea were routinely swapping genetic material with other species - often across huge taxonomic distances". " 'There's promiscuous exchange of genetic information across diverse groups,' says Michael Rose, an evolutionary biologist at the University of California, Irvine." "As early as 1993, some were proposing that for bacteria and archaea the tree of life was more like a web. In 1999, Doolittle made the provocative claim that 'the history of life cannot properly be represented as a tree'.13 'The tree of life is not something that exists in nature, it's a way that humans classify nature,' he says."
"Recent research suggests that the evolution of animals and plants isn't exactly tree-like either." "A team at the University of Texas at Arlington found a peculiar chunk of DNA in the genomes of eight animals - the mouse, rat, bushbaby, little brown bat, tenrec, opossum, anole lizard and African clawed frog - but not in 25 others, including humans, elephants, chickens and fish. This patchy distribution suggests that the sequence must have entered each genome independently by horizontal transfer."34 "HGT [horizontal gene transfer] has been documented in insects, fish and plants, and a few years ago a piece of snake DNA was found in cows." "Biologist Michael Syvanen of the University of California, Davis... compared 2000 genes that are common to humans, frogs, sea squirts, sea urchins, fruit flies and nematodes. In theory, he should have been able to use the gene sequences to construct an evolutionary tree showing the relationships between the six animals. He failed."
"The problem was that different genes told contradictory evolutionary stories." " 'We've just annihilated the tree of life. It's not a tree any more, it's a different topology [design or shape] entirely,' says Syvanen." "It is clear that the Darwinian tree is no longer an adequate description of how evolution in general works." "Rose goes even further. 'The tree of life is politely buried, we all know that,' he says. 'What's less accepted is that our whole fundamental view of biology needs to change.' Biology is vastly more complex than we thought, he says." " 'The tree of life was useful,' says Bapteste. 'It helped us to understand that evolution was real. But now we know more about evolution, it's time to move on.' "26
Evolutionists write: "The meaning, role in biology, and support in evidence of the universal 'Tree of Life' (TOL) are currently in dispute. Some evolutionists believe... that we can with available data and methods reconstruct this tree quite accurately, and that we have in fact done so, at least for the major groups of organisms. Other evolutionists... do not doubt that some... branching tree can in principle represent the history of all life. Still other evolutionists, ourselves included, question even this most fundamental belief, that there is a single true tree." "Darwin claimed that a unique inclusively hierarchical pattern of relationships between all organisms based on their similarities and differences was a fact of nature." Yet "the only data sets from which we might construct a universal hierarchy including prokaryotes, the sequences of genes, often disagree and can seldom be proven to agree. Hierarchical structure can always be imposed on or extracted from such data sets by algorithms designed to do so, but at its base the universal TOL rests on an unproven assumption about pattern that, given what we know about process, is unlikely to be broadly true." There is "the possibility that hierarchy is imposed by us rather than already being there in the data."12 "The finding that, on average, only 0.1% to 1% of each genome fits the metaphor of a tree of life overwhelmingly supports the... argument that a single bifurcating tree is an insufficient model to describe the microbial evolutionary process." "When chemists or physicists find that a given null hypothesis can account for only 1% of their data, they immediately start searching for a better hypothesis. Not so with microbial evolution, it seems, which is rather worrying. Could it be that many biologists have their heart set on finding a tree of life, regardless of what the data actually say?"10 "A single, uninterrupted TOL does not exist, although the evolution of large divisions of life for extended time intervals can be adequately described by trees." "Tree topology tends to differ for different genes."26 The genomes of all life forms are collections of genes with diverse evolutionary histories." "The TOL concept must be substantially revised or abandoned because a single tree topology or even congruent topologies of trees for several highly conserved genes cannot possibly represent the history of all or even the majority of the genes." "The 'strong' TOL hypothesis, namely, the existence of a 'species tree' for the entire history of cellular life, is falsified by the results of comparative genomics." "So the TOL becomes a network, or perhaps most appropriately, the Forest of Life that consists of trees, bushes, thickets..., and of course, numerous dead trunks and branches."22
Kevin Peterson, a molecular paleobiologist at Dartmouth College, "has been reshaping phylogenetic trees for the past few years, ever since he pioneered a technique that uses short molecules called microRNAs to work out evolutionary branchings. He has now sketched out a radically different diagram for mammals: one that aligns humans more closely with elephants than with rodents."
" 'I've looked at thousands of microRNAs, and I can't find a single example that would support the traditional tree,' he says. The technique "just changes everything about our understanding of mammal evolution'."
"And as he continues to look, he keeps uncovering problems, from the base of the animal tree all the way up to its crown."
"Peterson and his team are now going back to mammalian genomes to investigate why DNA and microRNAs give such different evolutionary trajectories. 'What we know at this stage is that we do have a very serious incongruence,' says Davide Pisani, a phylogeneticist at the National University of Ireland in Maynooth". --Dolgin, Elie. 28 June 2012. Rewriting Evolution. Nature, Vol.486, pp.460-462.
This is huge. Professional evolutionists spend most of their time adjusting their "tree of life". They have fun thinking how one type of creature "developed" into another type, how abilities "arose" or "emerged" here and there, but that is just playing at science. These articles show that, while they still cling to their belief in evolution, the truth is becoming inescapable to a few evolutionists who dare to look at the facts: Darwin was wrong; microbes, insects, plants, and animals do not fit a "tree of life" with linear descent. There is no pattern to their similarities and differences because each one is a uniquely designed, complete creature.
The big fudge
There
were warnings of problems in the very "tree of life" diagrams
evolutionists were fabricating. If you believed these diagrams
of new forms arising from old forms, then you believed that parts can be lost from a creature
and then exactly re-invented in the same creature millions of years
later. And that might happen several times! Not only
that, the same part might appear independently in unrelated creatures. For example, wings
would have had to evolve completely independently four times:
in insects, flying reptiles, birds, and bats.
|
|
|
When a creature produces light with chemicals it is called bioluminescence. "A remarkable diversity of marine animals and microbes are able to produce their own light, and in most of the volume of the ocean, bioluminescence is the primary source of light." "On land, fireflies are the most conspicuous examples, but other luminous taxa include other beetles, insects like flies and springtails, fungi, centipedes and millipedes, a snail, and earthworms." Evolutionists think bioluminescence evolved independently 40 to 50 separate times! --Haddock, Steven H.D., Mark A. Moline, James F. Case. 2010. Bioluminescence in the Sea. Annual Review of Marine Science, Vol. 2, pp. 443-493. |
In another study, evolutionists concluded that the cecal appendix evolved independently at least 32 separate times in mammals.41 This dodge is quite common in "evolutionary science".
Instead of heeding the warnings and scrapping the diagrams, they solved the problem by giving it a name: either "convergent" or "parallel" evolution (according to the situation), and it became standard in evolutionist writings. Now they just casually throw out statements like "convergent evolution is widespread across the mammal tree of life". --Helgen, Kristofer M. 28 October 2011. The Mammal Family Tree. Science, Vol. 334, pp. 458-459.
"Convergent evolution" is not a scientific explanation. It is a rationalization designed to explain away some of the major problems with the conventional evolutionary theory of divergence into a branching pattern that would be expected from common descent via natural selection. Creationists and Intelligent Design theorists account for the similarities as the result of engineering, using a common design that works. - Jerry BergmanIn their sales pitch to the public, evolutionists use the gimmick, "if a million monkeys typed randomly for millions of years, eventually one would type one of Shakespeare's sonnets", and people think "well, maybe...". How about the monkeys typing the same sonnet twice or more? It would be shocking if random mutation-natural selection produced any working part, let alone the same part twice or more from scratch. With convergent/parallel evolution, evolutionists could fudge any situation.
Perhaps the most brazen example of this can be seen in the "parallel evolution" of two types of mammals, placental (such as humans) and marsupial (such as kangaroos). Evolutionists tell us that each group evolved separately, yet many are remarkably similar, including cats, mice, wolves, moles, flying squirrels, anteaters, and others. This is whole animal duplication, not just an individual part.
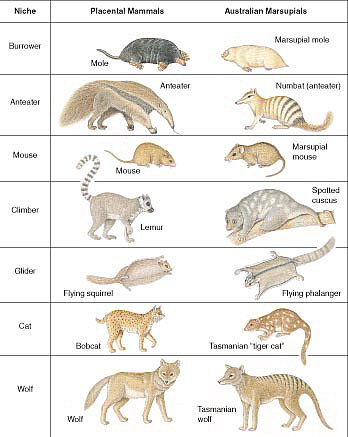
A normal person would be embarrassed if their theory of random change made such claims, but you cannot embarrass a fanatic. The only reason for the "convergent/parallel evolution" trick is to force the "tree of life" framework onto a world of uniquely designed creatures.
|
|
|
Animal genomes have billions of DNA base pairs, but each species is so unique that a very short stretch, or "barcode" is all you need to identify them. "For animals, the standard barcode is a 648 base pairs fragment of the mitochondrial gene cytochrome c oxidase 1 (COI). The use of COI for species identification and discovery has been extremely successful for the animal kingdom, and the BARCODE OF LIFE DATASYSTEMS database (BOLD) now contains more than 4.2 million validated barcodes." --Coissac, Eric, Peter M. Hollingsworth, Sebastien Lavergne, Pierre Taberlet. 2016. From barcodes to genomes: extending the concept of DNA barcoding. Molecular Ecology, Vol. 25, pp. 14231428. |
Design
flaws? 
Evolutionists say that while no intelligent designer would
design anything with flaws, evolution is a mechanical process of trial and
error, so evolution easily explains the existence of flaws in biology. In his 1986 book, "The Blind Watchmaker," the
famous evolutionary biologist Richard Dawkins wrote about the eye, "Any engineer would
naturally assume that the photocells would point towards the light, with their
wires leading backwards towards the brain.
He would laugh at any suggestion that the photocells might point away,
from the light, with their wires departing on the side nearest the light. Yet this is exactly what happens in all
vertebrate retinas." "This means that
the light, instead of being granted an unrestricted passage to the photocells,
has to pass through a forest of connecting wires, presumably suffering at least
some attenuation and distortion".
Evolutionist professors have told this to their students for years,
while looking at them with perfect vision.
It is true that "images projected onto the retina have to pass several layers of randomly oriented and irregularly shaped cells with intrinsic scatterers before they reach the light-detecting photoreceptor cells". "When light passes through multiple layers of cells, as in tissues, images rapidly deteriorate".16
"The mammalian retina and the peripheral retina of humans and primates are organized in a seemingly reverse order with respect to the light path. This arrangement places the photoreceptors, responsible for light absorption, as the last cells in the path of light, rather than the first. Therefore, the incident light must propagate through five reflecting and scattering layers of cell bodies and neural processes before reaching the photoreceptors. This 'inverted' retinal structure is expected to cause blurring of the image and reduction in the photon flux reaching the photoreceptors, thus reducing their sensitivity."25
Then researchers took a closer look at glial cells in the eye. "Glial cells are the most abundant cell types in the central nervous system. They surround neurons to support and insulate them." "Muller cells, the major type of glial cells in the retina, are responsible for the homeostatic and metabolic support of retinal neurons. By mediating transcellular ion, water, and bicarbonate transport, Muller cells control the composition of the extracellular space fluid. Muller cells provide trophic and anti-oxidative support of photoreceptors and neurons and regulate the tightness of the blood-retinal barrier. By the uptake of glutamate, Muller cells are more directly involved in the regulation of the synaptic activity in the inner retina. This review gives a survey of recently discoved new functions of Muller cells. Muller cells are living optical fibers that guide light through the inner retinal tissue."37
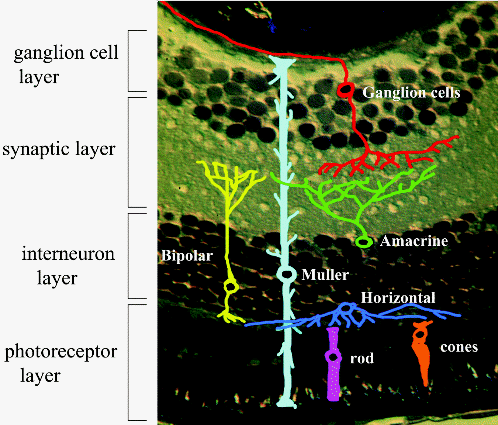
Muller cells span the entire thickness of the retina. "Individual Muller cells act as optical fibers. Furthermore, their parallel array in the retina is reminiscent of fiberoptic plates used for low-distortion image transfer. Thus, Muller cells seem to mediate image transfer through the vertebrate retina with minimal distortion and low loss."16
"The basic fiberoptic plate-like structure is especially characteristic for the retinae of all mammals with the exception of the fovea centralis of humans and higher primates, the region of our retina that is responsible for sharp vision; here, the photoreceptor cells are not obscured by any inner retinal layers at all."16
"Every mammalian Muller cell is coupled to one cone photoreceptor cell (responsible for sharp seeing under daylight conditions, i.e., photopic vision) plus a species-specific number of rod photoreceptor cells (about 10 in both man and guinea pig), serving low light level (scotopic) vision."16
"Light of relevant wavelengths for cone visual pigments is directed towards the cones, while light of wavelengths more suitable for rod vision is allowed to leak outside the Muller cells towards the surrounding rods."25
"The fundamental features of the array of glial cells are revealed as an optimal structure designed for preserving the acuity of images in the human retina. It plays a crucial role in vision quality, in humans and in other species."24
As usual, once we learn more about something in biology, evolutionist claims fall apart.
Reliving evolution?
An old evolution myth still
hanging around is the notion that things that look like gill-slits, tails,
etc. in developing human embryos show the embryo repeating
all the stages of evolution. In 1866, Ernst Haeckel
proposed his "biogenetic law" (not to be confused with
the law of biogenesis that says life only comes from life). His
idea was that growing vertebrate embryos went through all the forms
of their supposed evolutionary ancestors ("ontogeny recapitulates
phylogeny"). He published drawings comparing growing
embryos of a number of animals such as the pig, cat, salamander,
etc. to growing human embryos. The similarities that he said
he found helped persuade people to believe the theory of evolution.
Scientists eventually discovered enough about embryology to
quietly discard the "biogenetic law", but it was not until
a careful photographic study of growing vertebrate embryos was conducted
in 1997 that Haeckel's deceit was fully revealed. They
found that his drawings were so far from reality that they could
not have been done from the actual embryos.38 He
must have faked them.
Vanishing
vestigials
If
evolutionists do not know what something does, they assume it is
useless, as we will see with "junk DNA". One of
their "proofs" of evolution
has been that as creatures evolve, some body parts that were useful
long ago become less important in the new and improved creatures.
Eventually these parts no longer function and they shrink in
size. Evolutionists called them "vestigial organs".
In the late 1800s they made long lists of vestigial organs in humans, including the tonsils, pineal
gland, thymus, and appendix.5
In the years since, advances in our understanding of anatomy and biology have
knocked them off the lists one by one. Yet the notion lingers
on that "there is something to it".
|
|
|
Over 70% of all primate and rodent taxonomic families contain species with an appendix.40,41 In 2009, researchers at Arizona State and Duke Universities reported that the little appendix is a "safe house" for important gut bacteria. If the intestine becomes infected and is forced to flush everything out (diarrhea), the good bacteria stored in the appendix are there to return the intestine to normal working order. "Maybe it's time to correct the textbooks," says William Parker, Ph.D., assistant professor of surgical sciences at Duke and the senior author of the study. "Many biology texts today still refer to the appendix as a vestigial organ."14 |
The vestigial organ idea helped fool millions of people into believing the theory of evolution. Today, this "proof" is down the toilet.
Forget about
neutral mutations
So-called
nonsynonymous mutations change protein sequences, and
they are damaging. "Synonymous mutations in protein-coding genes do not alter
protein sequences and are thus generally presumed to be neutral or nearly
neutral."
Researchers "constructed 8,341 yeast mutants each carrying a synonymous, nonsynonymous or nonsense mutation in one of 21 endogenous genes with diverse functions and expression levels and measured their fitness".
"Three-quarters of synonymous mutations resulted in a significant reduction in fitness, and the distribution of fitness effects was overall similaralbeit nonidenticalbetween synonymous and nonsynonymous mutations. Both synonymous and nonsynonymous mutations frequently disturbed the level of mRNA expression of the mutated gene".
" under any environment, most synonymous mutations are strongly non-neutral, and the distribution of fitness effects of synonymous and nonsynonymous mutations are overall similar. There is no particular reason why our results would not generalize to other organisms".
"Our results also imply that synonymous mutations are nearly as important as nonsynonymous mutations in causing disease".
"Because many biological conclusions rely on the presumption that synonymous mutations are neutral or nearly neutral, its invalidation has broad implications." - Shen, Xukang, Siliang Song, Chuan L, Jianzhi Zhang. 8 June 2022. Synonymous mutations in representative yeast genes are mostly strongly non-neutral. Nature. DOI:10.1038/s41586-022-04823-w
Violating
the law
The theory of Evolution violates two
laws of science. The Second Law of Thermodynamics (law of
increasing entropy) says that concentrations of things spread out over time.
If you heat one room in a house, then open the door to that room,
eventually the temperature in the whole house evens out (reaches equilibrium).
Knowing how far this evening-out has progressed at any point in time
tells you the entropy. Entropy can measure the loss of a system's ability
to do work. Entropy is also a measure of disorder:
An isolated system or a system in a uniform environment (which for the present consideration we do best to include as the part of the system we contemplate) increases its entropy and more or less rapidly approaches the inert state of maximum entropy. We now recognize this fundamental law of physics to be just the natural tendency of things to approach the chaotic state (the same tendency that the books of a library or the piles of papers and manuscripts on a writing desk display) unless we obviate it. Famous physicist Erwin Schrödinger from his book What Is Life?
And that is where evolution theory hits an impenetrable wall. Natural processes proceed in only one direction, toward equilibrium and disorder. Things fall apart over time, they do not get more organized. We can overcome this by making a machine and adding energy, but the Second Law prevents such a machine from assembling spontaneously from raw materials.
The Law of Biogenesis was established by Louis Pasteur three years after Darwin's book was published, and simply says that life only comes from life. Living cells divide to make new cells, and fertilized eggs and seeds develop into animals and plants, but chemicals never fall together and life appears. Evolutionists often call certain chemicals "the building blocks of life", giving people the false impression that you just stack the building blocks together and you get life. No one has ever done that, including the famous 1953 Miller/Urey experiment where all they got were clumps of amino acids. Many people mistakenly think scientists have made life from chemicals in the lab, but they have not (though many have tried very hard). If one were to succeed, you would know about it. He would be treated like a superhero, so desperate are evolutionists on this matter.For something to be a law of science, it can never be found to have been violated, even once, over thousands of trials. No exceptions. A theory that violates two laws of science is in big trouble.
When confronted with the Second Law of Thermodynamics, evolutionists usually use two tricks to try to escape. The first is to state that "it only applies to closed systems, and biological creatures are open systems, so it doesn't affect evolution" (they actually intend to say isolated, not closed, but we know what they mean). The fact is that the Second Law applies to all systems, open or closed, and to all actions and chemical reactions, from molecules to galaxies. The words "except for..." are not in this universal law. A thermodynamics system is simply any part of the universe we want to study. If we are doing an experiment in a bottle, the inside of the bottle is our system and the bottle itself is the "walls" of the system. There are only 3 kinds of systems: if no energy or matter can pass through the walls, it is an isolated system; if energy can pass through but matter cannot, it is a closed system; if both energy and matter can pass through the walls, it is an open system. Now, it is true that the laws of thermodynamics and entropy are defined in terms of isolated systems, because that is the simplest way to express them. However, experts who write textbooks on the subject are quick to say that isolated systems do not occur in nature. For practical applications, a procedure called the Legendre Transform mathematically converts entropy to a variable called Gibbs free energy that is useful for working with real-world systems. Most natural systems are open, but it is convenient to model them as closed. For example, even though a bacterium is an open system, modeling it as a closed system makes it easier to understand chemical reactions in it.2,8
You are an open system. You eat food (which comes from outside yourself) and your body survives and grows. Evolutionists believe that all we need is an open system with sufficient energy flowing into it for evolution to succeed. If that were so, you could just stand right behind a jet engine as the aircraft prepares for takeoff, absorb that blast of energy, and evolve to a higher life form. In reality, of course, you would be incinerated because absorbing energy without a mechanism to convert it to a useful form and employ it is destructive or useless. The mechanism must be very specific. Sticking food in your ear will not work; it must go into your mouth and through the digestive system. And the mechanism must be in place and functioning first, before energy is added, or the energy is wasted. The "closed system" ploy is just an attempt to avoid dealing with the Second Law because the Law prohibits any functioning biological mechanism from falling together by pure chance, without assistance or plan, using only the properties of matter. Evolutionists also believe that chemical evolution could have started when a high-energy spark, like lightening, split molecules into radicals and ions that randomly combined with each other to produce the new, highly complex molecules their theory needs. They ignore the fact that, following the Second Law, it would also produce all other possible combinations of molecules, and many of these chemicals would work against chemical evolution. Without a sufficient concentration of the pure chemicals needed, with the proper chirality and ratios to each other, the main result would be a useless tar like what the famous Miller/Urey experiment produced in 1953.
The second trick they use is to say that "when you freeze water, the disordered molecules become beautifully ordered ice crystals or snowflakes. If water can bypass the Second Law and organize its molecules by a natural process, why not the chemicals of life?" At room temperature, water molecules are bouncing off each other and you have water. When you take away heat and they freeze, water molecules stick to each other with weak molecular bonds, forming ice crystals and snowflakes because of the shape of the H2O molecule. The same thing happens if you put a bunch of weak magnets in a jar and shake it. The magnets bounce around. When you stop, the magnets stick together. They are at a lower energy level. There is order, yet no complexity - just a simple repetitive structure that does not violate the Second Law.
But guess what. Amino acid molecules that form proteins, and nucleotide molecules that form DNA and RNA resist combining at any temperature. To combine, they need the help of mechanisms in a living cell or a biochemist in an organic chemistry laboratory.18 It means that nothing happens in the primeval soup, the pond of chemicals where evolutionists believe life began.
DNA is made of only right-handed versions of nucleotides, while proteins are made of only left-handed versions of amino acids. Yet any random chemical reaction that produced nucleotides or amino acids would make an equal mix of left and right-handed versions of each. Even if the thousands of nucleotides needed to form a DNA molecule, or the hundreds of amino acids needed to form a protein molecule were able to combine from the mix, they would be a jumble of left and right-handed versions that could not function at all. This is the problem of "chirality", and evolutionists have never been able to solve it.
Ilya Prigogene coauthored a paper in 1972 that says in an open "system there exists a possibility for formation of ordered, low-entropy structures at sufficiently low temperatures. This ordering principle is responsible for the appearance of ordered structures such as crystals... Unfortunately this principle cannot explain the formation of biological structures."36 Prigogene won the Nobel Prize in Chemistry in 1977 for research on dissipative structures, such as tornados, for contributions to nonequilibrium thermodynamics, and for bridging the gap between biology and other sciences. Evolutionists wrongly claim he won for showing how thermodynamics could explain the formation of organized systems, from fluctuations in chaos, that lead to the origin of life. They thought he was their hero. Over thirty-five years later, nothing has come of it. There is no escape from the Second Law of Thermodynamics. It prohibits the spontaneous origin of life and macroevolution.
Mindboggling complexity cells and proteinsEven a single cell is not simple. In Darwin's day researchers looked at cells under the microscope and saw little balloons filled with goo they called protoplasm, so they thought cells were simple forms of life. Over 160 years later we know that there are many types of cells, and each cell is a little city at work.
Forty-three biologists introduced their Cell Atlas in the journal Science in May 2017, a "subcellular map of the human proteome". "Cells are internally organized into compartments called organelles. The spatial partitioning provided by organelles creates an enclosed environment or surface for chemical reactions tailored to fulfill specific functions. These functions are tightly linked to a specific set of proteins."
They made this atlas to organize the many thousands of proteins in human cells, and have categories for new ones as they are discovered. This is Darwin's simple goo.
Thul,
Peter J. et al., 26 May 2017, A Subcellular Map of the Human Proteome,
Science, Vol. 356, No. 6340, 12 pages, DOI:10.1126/science.aal3321
Check out one small part of a cell - the pores that let things pass in and out of a nucleus; the Nuclear Pore Complex.
The smallest known genome (Mycoplasma genitalium) has 482 genes.19 The minimum possible for an organism to survive is probably 200 to 300 genes. Most bacteria have 1000 to 4000 genes. Everything about the cell is stunningly complex. A popular textbook on the cell1 is 1600 pages long and weighs 7 pounds. Plants and animals contain a great variety of cells; the human body has about 210 different types.
Cells are made of proteins, and everything that goes on in a creature involves proteins interacting with each other. See a brilliant animation of protein construction in this short clip from an Illustra Media video, with Dean Kenyon's comment.
Proteins are generally 50 to 2000 amino acids long; a typical one has about 300 amino acids.1 Ribosomes are molecular machines that build proteins in cells, using messenger RNA as the template. Here is an overview of how a bacterial ribosome "translates" RNA into protein. Every protein in bacteria is made this way.
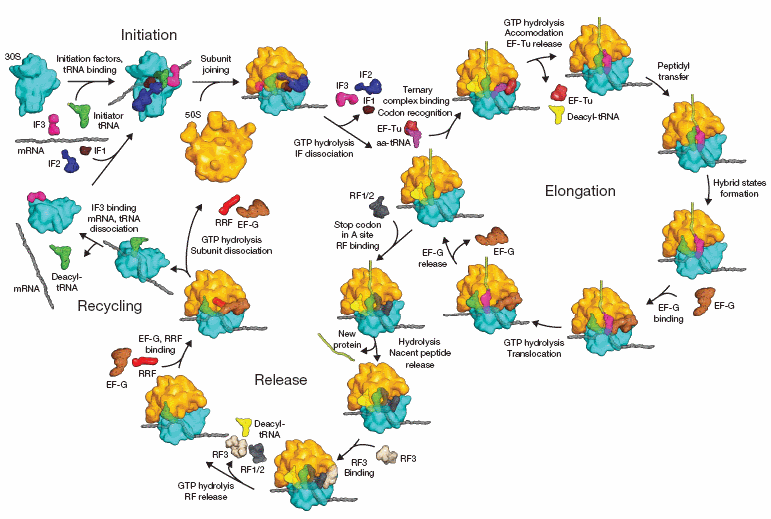
From:
Schmeing, T. Martin, V. Ramakrishnan. 29 October 2009. What recent
ribosome structures have revealed about the mechanism of translation.
Nature, Vol. 461, pp. 1234-1242.
A protein is not just a long ribbon of amino acids strung together from the DNA pattern. It folds itself into a 3D structure.

|
Diagram of a folded protein |
Origami |
|
|
"Inside every cell in your body, billions of tiny molecular machines are hard at work. They're what allow your eyes to detect light, your neurons to fire, and the 'instructions' in your DNA to be read, which make you the unique person you are. These exquisite, intricate machines are proteins. They underpin not just the biological processes in your body but every biological process in every living thing. |
Currently, there are around 200 million known proteins, with another 30 million found every year. Each one has a unique 3D shape that determines how it works and what it does.
If you could unravel a protein you would see that it's like a string of beads made of a sequence of different chemicals known as amino acids. These sequences are assembled according to the genetic instructions of an organism's DNA.
Attraction and repulsion between the 20 different types of amino acids cause the string to fold in a feat of 'spontaneous origami', forming the intricate curls, loops, and pleats of a protein's 3D structure.
For decades, scientists have been trying to find a method to reliably determine a protein's structure just from its sequence of amino acids. This grand scientific challenge is known as the protein folding problem.
We started working on this challenge in 2016 and have since created an Artificial Intelligence (AI) system known as AlphaFold. Our latest version can now make accurate predictions of what shape a protein will form based on its sequence of amino acids." - From https://deepmind.com/research/case-studies/alphafold
"DeepMind [owner of AlphaFold] is an AI company that was acquired by Google in 2014. DeepMind trained its system using 128 specialized processors for a couple of weeks; it now returns potential structures within a couple of days." - https://arstechnica.com/science/2020/11/deepmind-ai-handles-protein-folding-which-humbled-previous-software/
Researchers have had to use artificial intelligence computers to translate DNA code into folded structures. What kind of intelligence wrote the linear codes for proteins into DNA?
The temperature and chemical concentrations must be right for it to fold correctly, and many proteins get help from special proteins called "molecular chaperones". Chaperones can keep proteins separated from each other while they are folding, prevent mistakes in folding, and even unfold mistakes to give the protein a second chance to get it right. After helping one protein fold, a chaperone will go help another one fold.
"A
chaperone protein (bottom, yellow) called SecB guides the folding
of another protein (transparent) in this artist's illustration."--Science News, December 1, 2007, Vol. 172, p. 342
Making and folding proteins goes on continuously throughout the body. Misfolding can lead to more than proteins that don't work. In humans, bunches of them (aggregates) can lead to diseases such as Alzheimer's, Huntington's, or sickle cell. "Proteins are so precisely built that the change of even a few atoms in one amino acid can sometimes disrupt the structure of the whole molecule so severely that all function is lost."1 All proteins stick (bind) to other molecules. But each can bind to only a few of the thousands it encounters. "An average protein in a human cell may interact with somewhere between 5 and 15 different partners."1 Their shapes fit each other like a hand in a glove. "Proteins can form enormously sophisticated chemical devices." "The most impressive tasks are carried out by large protein assemblies formed from many protein molecules." "Each of the central processes in a cell... is catalyzed by a highly coordinated, linked set of 10 or more proteins."1 The parts of a cell where proteins are made (ribosomes) are themselves made of many different proteins. "The complexity of living organisms is staggering."1 In the face of this breathtaking complexity, evolutionists have tried to find the basic things necessary for a cell to function. So far they have found 17 general categories1:
- Replication, recombination, and repair
- Transcription
- Cell cycle control, mitosis, and meiosis
- Defense mechanisms
- Cell wall/membrane biogenesis
- Signal transduction mechanisms
- Intracellular trafficking and secretion
- Translation
- Post-translational modification, protein turnover, chaperones
- Energy production and conversion
- Carbohydrate transport and metabolism
- Amino acid transport and metabolism
- Nucleotide transport and metabolism
- Coenzyme transport and metabolism
- Lipid transport and metabolism
- Inorganic ion transport and metabolism
- Secondary metabolite biosynthesis, transport, and catabolism
Each category requires many proteins. All have to be in place and working together or the cell is wrecked.
Scientists have found that the number of genes a creature has is not a good measure of how complex it is. For example, the human genome is 23 times larger than the fruit fly genome (3.2 billion base pairs versus 137 million), yet humans have less than twice the number of protein coding genes (21,000 versus 13,000) . Yeast has about 6,000 genes.
|
|
|
The tiny water flea
Daphnia pulex has more genes than humans do;
up to 39,000 at last count. |
|
|
|
|
|
|
|
So does the pea aphid
Acyrthosiphon pisum,
with 34,600. |
The main reason for biological complexity must be due to something else. "Alternative splicing" is an important part of it.
"Humans do not have many more genes than mice, fruit flies, or worms. This observation raises the question of how humans can be so much more morphologically and behaviorally complex than these other metazoans."
|
|
|
"One of the most unanticipated findings in molecular biology was the discovery that eukaryotic genes are discontinuous, with protein-coding segments or exons disrupted by noncoding segments or introns." Splicing exons together in different ways allows a single gene to code for multiple proteins. |
"The average human gene contains eight exons and seven introns, producing an average of three or more alternatively spliced mRNA isoforms. 100% of human genes produce at least two alternative mRNA isoforms."
"It has become apparent that precursor messenger RNA (pre-mRNA) splicing can occur to a great extent that scales with organismal complexity. Indeed, although the mouse and human genomes contain similar numbers of genes, alternative pre-mRNA splicing occurs in >95 to 100% of human genes, compared with ~63% of mouse genes. Thus, one function of alternative splicing is to significantly expand the form and function of the human proteome. Alternative splicing can serve many regulatory functions."
|
|
|
"Intron removal in protein-coding mRNAs (and long noncoding RNAs) occurs in a large ribonucleoprotein (RNP) machine called the spliceosome." |
"The spliceosome functions in a complex and dynamic assembly, reaction, and disassembly cycle in which five small nuclear ribonucleoprotein (snRNP) complexes (U1, U2, U4/U6, and U5) recognize and assemble on each intron to ultimately form a catalytically active spliceosome. Spliceosome assembly needs to occur every time an intron is removed from a pre-mRNA in a eukaryotic nucleus. The protein composition of the spliceosome dynamically changes as the assembly and subsequent catalytic steps occur."
"Alternative splicing may be a key aspect related to the phenotypic complexity of Homo sapiens."-- Lee, Yeon, Donald C. Rio. 2015. Mechanisms and Regulation of Alternative Pre-mRNA Splicing. Annual Review of Biochemistry, Vol. 84, pp. 291-323.The old view of proteins was that "each gene encodes a single protein, and each protein serves a single function. However, this orderly view of biology has turned out to be overly simplistic. Single genes can encode different proteins as a result of alternative splicing, and single proteins often serve multiple functions."
|
|
|
"Moonlighting - the performance of more than one function by a single protein - is becoming recognized as a common phenomenon". "Many proteins have more than one moonlighting function. In some cases, different organisms have recruited the same protein to serve different moonlighting functions. In other cases, a protein serves several moonlighting functions within the same organism, and even within the same cell type." |
"Moonlighting functions are provided by other parts of the protein. Since the region devoted to the canonical [main] function of a protein occupies only part of the molecule, extensive regions are available for binding to other macromolecules."
"Correct orchestration of moonlighting functions requires that proteins must be either produced in, or directed to, different places in the right quantities and at the right times."
"An inescapable conclusion based upon the torrent of discoveries of moonlighting proteins is that cellular physiology is more complex than we imagined based upon the comfortable reductionist viewpoint in which each protein serves a single function."-- Copley, Shelley D. 2012. Moonlighting is mainstream: Paradigm adjustment required. Bioessays Vol. 34, No. 7, pp. 578588.
So evolutionists have to believe that for each protein, pure chance laid out long strings of amino acids that fold themselves into the exact shapes needed to interact with other specialized proteins and, where needed, get help from chaperone proteins which themselves appeared by chance. The necessary proteins cannot be invented one at a time. Either they are all there, ready to work together, or nothing happens and they disintegrate. Yet even if it could design proteins, mutation-natural selection would only work on one at a time sporadically over many years. Considering just the complexity of proteins, the notion of creating them with mutation-natural selection is as silly as asking someone to build a television set with a spoon and a toothbrush. If Darwin had known what we have learned about proteins, he probably would have abandoned the theory of evolution.
Darwin himself wrote in chapter 6 of On the Origin of Species that "natural selection can act only by the preservation and accumulation of infinitesimally small inherited modifications, each profitable to the preserved being... If it could be demonstrated that any complex organ existed, which could not possibly have been formed by numerous, successive, slight modifications, my theory would absolutely break down."
Duplicate
genes
|
|
Do evolutionists admit defeat? Never! They temporarily isolate their DNA workshop from natural selection, saying the mutations in DNA for building a complicated new part accumulate quietly in duplicate genes. Then, millions of years later, all are in place. The new part starts working, natural selection chooses it, and the improved creature is off to the races. This scenario exists only in the mind of the evolutionist. |
It is taught in textbooks that gene duplication is the major way to drive evolution. Evolutionists believe that by mutation and natural selection, one or all copies of a duplicated gene eventually encode new proteins (a process they call neofunctionalization). Over millions of years, small simple genomes would thus evolve into large, complex ones, giving rise to all life forms.
This was thought to be the only mechanism to generate new genes from existing ones. However, biologists are now becoming more and more convinced theoretically and empirically that most duplicated gene copies undergo degenerative, rather than constructive, mutations, ending up in nonfunctionalization. Faced with this dilemma, evolutionists have thought of ways to theoretically keep duplicates functional. But they still depend on mutation and natural selection for neofunctionalization.
Polyploidy refers to an increase in the number of sets of chromosomes per cell. It may arise naturally when a cell fails to divide after DNA replication. If the cell with doubled genome is involved in the generation of sex cells (meiosis), polyploid organisms may be subsequently produced upon fertilization.
In reality, the first event awaiting a duplicated gene is silencing. The best studied mechanism of silencing is through methylation of cytosine bases in CG islands around promoters. After that, methylated cytosines tend to spontaneously lose amino acids and are substituted with thymine bases. The phenomenon is known as CG depletion. Duplicated genes are especially prone to CG depletion. Silenced duplicates may also undergo other mutations. Indeed, extensive genomic change can be detected within a few generations after synthetic polyploidy. Duplicated genes are lost exponentially with time and are nonfunctionalized by the time silent sites have diverged by only a few percent.
The bottom line is that gene duplications are aberrations of cell division processes and are more likely to cause malformation or diseases rather than selective advantage. Plants can tolerate duplications, especially polyploidy, better than animals because of differences in the way they reproduce. To maintain genomic stability, all cells have built-in mechanisms to silence duplicated genes, after which they fall victim to degenerative mutations.
Gene regulatory sequences and hierarchies determine complexity. Regulation of supposedly duplicated gene clusters and gene families is irreducibly complex, and demands the simultaneous development of fully functional multiple genes and switching networks, contrary to Darwinian gradualism. This constitutes an insurmountable barrier for the theory. - Liu, Yingguang, Dan Moran. August 2006. Do new functions arise by gene duplication? Journal of Creation, Vol. 20, No. 2, pp. 82-89. https://creation.com/do-new-functions-arise-by-gene-duplicationEvolutionary scientists know they need to explain the origin of genetic information. However, instead of discussing new information, they tend to focus on new genes. These are sometimes known as de novo genes. While evolutionists have proposed a number of mechanisms to generate new information, none of them do what is claimed. Instead, they either break the genome or rearrange existing information. Even if the mechanisms did not cause disease, simply creating new sequences is not enough. The new sequences must be able to be read and not create mutations, which are almost exclusively deleterious. Even if the required genetic sequences could be generated, a significant number of beneficial mutations would be required to create new functional information. Evolution simply lacks the mechanism required to create new information. - Sanders, Harry F. III. January 30, 2021. New Genetic Information Proposals Fail. Answers in Depth, Vol. 16. https://answersingenesis.org/genetics/new-genetic-information-proposals-fail/
A review of many experiments with duplicate genes found that "duplication can and does facilitate important adaptations by tinkering with existing compounds". The "random mutations and recombinations considered were observed to tweak, tinker, copy, cut, divide, and shuffle existing genetic information around, but fell short of generating genuinely distinct and entirely novel functionality." "Gene duplication results in the copying and preservation of biological information, and not its transformation as something original." It is "insufficient in explaining the origination of the highly complex information pertinent to the essential functioning of living organisms." - Bozorgmehr, Joseph Esfandiar Hannon. 22 December 2010. Is Gene Duplication a Viable Explanation for the Origination of Biological Information and Complexity? Complexity, published online, DOI 10.1002/cplx.20365, pp. 1-14.
Repairing
mutations
So duplicate genes are not macroevolution's secret
laboratory. Furthermore,
everyone agrees that harmful mutations appear many, many times more
often than mutations needed for new construction ever could.
Over those millions of years, slightly harmful mutations
that are hidden, or not destructive enough for natural selection
to remove, would also quietly accumulate. This would produce
creatures loaded up with highly polluted genes. Survival of
the barely functional? We do not find this either because
cells have mechanisms that maintain the original
design of a creature within its variation
boundaries, and minimize the accumulation of mutations. These
include:
- A proofreading system that catches almost all errors
- A mismatch repair system to back up the proofreading system
- Photoreactivation (light repair)
- Removal of methyl or ethyl groups by O6 - methylguanine methyltransferase
- Base excision repair
- Nucleotide excision repair
- Double-strand DNA break repair
- Recombination repair
- Error-prone bypass46
Below is a description of the second type - mismatch of base pairs. How do you suppose this mechanism evolved randomly?
Stokstad,
Erik. 16 October 2015. DNA's repair tricks win chemistry's top prize.
Science, Vol. 350, No. 6258, p. 266
Harmful mutations happen constantly. Without repair mechanisms, life would be very short indeed and might not even get started because mutations often lead to disease, deformity, or death. So even the earliest, "simple" creatures in the evolutionist's primeval soup or tree of life would have needed a sophisticated repair system. But the mechanisms not only remove harmful mutations from DNA, they would also remove mutations that evolutionists believe build new parts. The evolutionist is stuck with imagining the evolution of mechanisms that prevent evolution, all the way back to the very origin of life.
Junk DNA
Only a small portion of a creature's
DNA is protein-coding genes (around 1.5% in humans). In the 1970s,
evolutionists began calling the rest of it "junk DNA",
saying this collection of useless evolutionary debris showed there
was no intelligent design involved. Decades later, researchers
are finding that the "junk" does vital
work. Some of this DNA plays a role in turning genes on and
off at the right moments in a developing embryo29. Other bits
separate coding and regulating sections, like punctuation marks
in writing, so that DNA is not a long run-on sentence30. Other
bits called Alu elements, found only in primates, can be spliced
in or out during RNA processing to make different versions of the
same gene.31
The "junk" label discouraged research into this part of the genome for many years; who would want to waste time studying it?
Then, in 2012, an article appeared in Science titled "ENCODE project writes eulogy for junk DNA". "This week, 30 research papers sound the death knell for the idea that our DNA is mostly littered with useless bases. A decade-long project, the Encyclopedia of DNA Elements (ENCODE), has found that 80% of the human genome serves some purpose". "The ENCODE effort has revealed that a gene's regulation is far more complex than previously thought, being influenced by multiple stretches of regulatory DNA located both near and far from the gene itself and by strands of RNA not translated into proteins, so-called noncoding RNA."-- Pennisi, Elizabeth. 7 September 2012. Science, Vol. 337, pp. 1159-1161.
"DNA repeated in tandem, once called junk DNA is an active part of the genome." "The genomes of eukaryotic species are made up of large amounts of repeated sequences. Among them, the most abundant fraction is constituted of satellite DNA (satDNA)". "Although originally satellite DNA was thought to be silent and inert, an increasing number of studies are providing evidence on its transcriptional activity supporting, on the contrary, an unexpected dynamicity." "Indeed, satellite DNA-derived transcripts play a structural function in heterochromatin formation and maintenance of both centromeres and telomeres, are involved in determining centromere identity interacting with CENP-A and kinetochore proteins, and control telomere length, capping and replication in a cell cycledependent manner." "These highly condensed structures are indispensable to preserve chromosome integrity and genome stability, preventing recombination events, and ensuring the correct chromosome pairing and segregation."-- Biscotti, Maria Assunta, Adriana Canapa, Mariko Forconi, Ettore Olmo, Marco Barucca. 2015. Transcription of tandemly repetitive DNA: functional roles. Chromosome Research, Vol. 23, pp. 463-477 DOI10.1007/s10577-015- 9494-4
"The days of "junk DNA" are over. When the senior authors of this article studied genetics at their respective universities, the common doctrine was that the nonprotein-coding part of eukaryotic genomes consists of interspersed, 'useless' sequences, often organized in repetitive elements". "This view has fundamentally changed".-- Stitz, Maria, Cristian Chaparro, Zhigang Lu, V. Janett Olzog, Christina E. Weinberg, Jochen Blom, 72 Alexander Goesmann, Christoph Grunau, Christoph G. Grevelding. 1 September 2021. Satellite-Like W-Elements: Repetitive, Transcribed, and Putative Mobile Genetic Factors with Potential Roles for Biology and Evolution of Schistosoma mansoni. Genome Biology and Evolution (GBE), Vol. 13, No. 10, pp. 1-20 DOI:10.1093/gbe/evab204
As usual, Evolution theory was misleading because it is pseudoscience.
Networks and
Systems Biology
The
living things of the world are extremely varied and intricately
made, yet the theory of evolution has always been about simplicity:
once upon a time, some chemicals assembled, began to make copies
of themselves, and little by little changed into all life forms.
Evolutionists like to use the words "simply" and
"merely" when telling their stories to the public. There
is certainly nothing complicated about the idea of mutation-natural
selection.
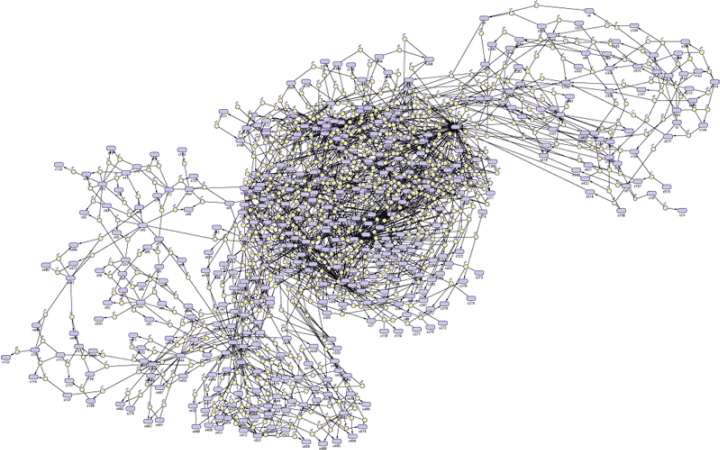

A small section of a biological system in an organism, displayed as
a 3D network
Networks and Systems Biology aims to understand the networks of interactions and effects of those interactions involving hundreds of different biological molecules simultaneously. - https://www.nature.com/subjects/networks-and-systems-biology
Systems biology has been responsible for some of the most important developments in the science of human
health and environmental sustainability. It is a holistic approach to deciphering the complexity of biological
systems that starts from the understanding that the networks that form the whole of living organisms are
more than the sum of their parts.
- Institute for Systems Biology
Discoveries in Systems Biology are the exact opposite of evolutionists' "merely, simple". Biological systems are vastly more complex than anyone could imagine. Some wonder if we will ever fully understand them.
"To make sense of the genome, systems biologists think in terms of networks. If two kinds of proteins or other biological molecules interact, they are connected on the network." "These network diagrams... show how individual pathways crisscross to form a tangled web. Each protein in a pathway can interact with molecules in other pathways, sometimes dozens of them." Additionally, "systems biologists produce complex maps of how genes and proteins interact, and these maps help scientists analyze results from drug screening." "Cells 'talk' to each other by passing chemical signals back and forth. They also sense their physical surroundings through proteins on their surfaces called integrins. All these cues serve to orient the cells in the body and inform them about how to behave so that they cooperate with the rest of the cells in the tissue." "The cells are not complete by themselves. They need signals from outside," says Mina J. Bissell of Lawrence Berkeley National Laboratory. "The unit of function literally is the tissue." - Patrick Barry. April 5, 2008. You, in a dish: cultured human cells could put lab animals out of work for chemical and drug testing. Science News, Vol. 173, No. 14, pp. 218-220.
"The interesting point coming out of all these studies is how complex these systems are; the different feedback loops and how they cross-regulate each other and adapt to perturbations are only just becoming apparent. The simple pathway models are a gross oversimplification of what is actually happening", says Mike Tyers, a systems biologist at the University of Edinburgh, UK. - Blow, Nathan. 16 July 2009. Untangling the protein web. Nature, Vol. 460, pp. 415-418.
"The life of every organism depends crucially on networks of interacting proteins that detect signals and generate appropriate responses. Examples include chemotaxis, heat shock response, sporulation, hormone secretion, and cell-cycle checkpoints.""When the information in molecular mechanisms that underlie the adaptive behaviour of living cells is laid out in graphical form, the molecular network looks strikingly similar to the wiring diagram of a modern electronic gadget. Instead of resistors, capacitors and transistors hooked together by wires, one sees genes, proteins and metabolites hooked together by chemical reactions and intermolecular interactions."

"Complex molecular networks, like electrical circuits, seem to be constructed from simpler modules: sets of interacting genes and proteins that carry out specific tasks and can be hooked together by standard linkages." - Tyson, John J., Katherine C. Chen, Bela Novak. 2003. Sniffers, buzzers, toggles and blinkers: dynamics of regulatory and signaling pathways in the cell. Current Opinion in Cell Biology, Vol. 15, No. 2, pp. 221231.
This is a map of how the genes in a cell of the budding yeast Saccharomyces cerevisiae interact with one another. Each color shows what a group of genes does. Genes in these functional networks interact with other genes throughout the cell; a cell of yeast.
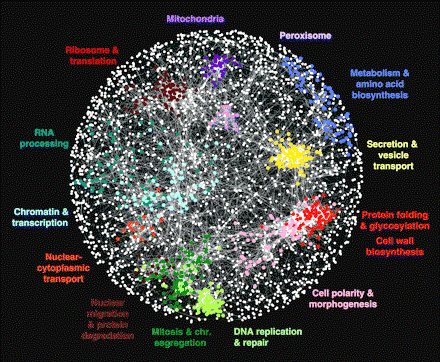
Costanzo,
Michael, et al. 22 January 2010. The Genetic Landscape of a Cell.
Science,
Vol. 327, No. 5964, pp. 425-431.
By 2010, real biologists had determined that gene regulatory networks (GRNs) build and operate all living things. There are gene regulatory networks for everything that happens in them, and some networks control other networks in a chain of command. Each species has a body plan, and it is encoded in the DNA. "Development of the body plan is caused by the operation of GRNs". "Embryonic development is an enormous informational transaction, in which DNA sequence data generate and guide the system-wide spatial deployment of specific cellular functions." That is, an embryo grows because GRNs tell other GRNs what to do at the right time and place and in the right order; it is tremendously complex. GRNs then guide the development of different types of cells, organs, and growth of the embryo into an adult. They also control each creature's abilities and the way it responds to changes around it. Among the most studied are sea urchins, which are low on the evolutionist's "tree of life".
An embryo has a particular growth program for the type of creature it will develop into. Yet it is likely that all the different programs are constructed from just a few types of sub-circuits. "Structurally similar sub-circuits, but composed of different regulatory genes" do "similar developmental jobs in different GRNs." This discovery gives researchers hope that they can use these "modules of developmental logic function" to decipher the "enormous mazes of interconnections in system level GRNs". You can tell what a sub-circuit does by its shape or structure. There is a sub-circuit for each task, and GRNs are made up of sub-circuits. The same control processes are used "throughout embryonic development because the problems that have to be solved are general: the initial spatial inputs have to be interpreted, the regulatory state then has to be locked down (the initial inputs are always transient), signals have then to be generated, other states have to be excluded, and differentiation drivers have to be activated. It is not surprising that all this requires a lot of sequential circuitry."
GRNs in embryos "are hierarchical in their overall structure. Their depth reflects the long sequence of regulatory steps required to complete any component of embryonic development." A GRN might have many layers of sub-circuits or very few, depending on its job. The last step in the chain of command is the signal for "batteries" (groups) of genes to change stem cells into specific types of cells (such as muscle, blood, nerve, etc.) at the right place. - Davidson, Eric H. 16 December 2010. Emerging properties of animal gene regulatory networks. Nature, Vol. 468, pp. 911-920.
Some evolutionists have publicly welcomed GRNs because a change in one controller can affect many genes, and we are back to simplicity. That is like saying a child can use Windows operating system on a computer. Just point and click with a mouse, and the computer does many complicated things. It is simplicity itself. So why are GRNs the death blow to evolution theory? It took computer and software engineers decades of intelligent design to build the computer and operating system. GRNs are the operating system of living things. The theory of evolution cannot explain how gene regulatory networks came to be. As with other insurmountable problems with the theory, this one remains in their pile marked "needs further study".
Today there is an explosion of knowledge going on in the study of gene regulatory networks. But it is not led, assisted, or even inspired by the theory of evolution. "We have little empirical knowledge on the evolutionary history of such networks." - Dean, Antony M., Joseph W. Thornton. September 2007. Mechanistic approaches to the study of evolution: the functional synthesis. Nature Reviews Genetics, Vol. 8, pp. 675-688.
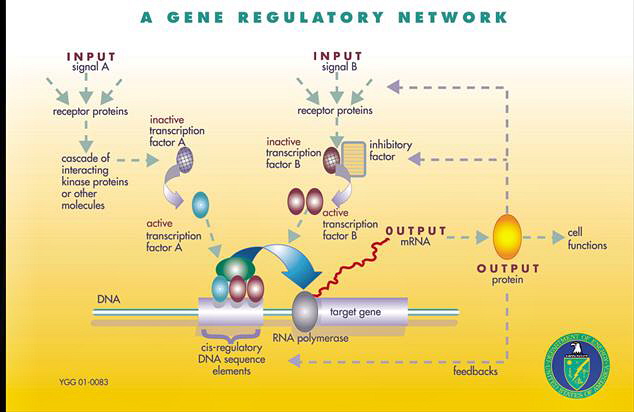
Some of the things GRNs have been found to do:
- Specialized GRNs determine which genes are active or inactive in each part of a developing creature
- GRN sub-circuits, usually consisting of 3 to 8 regulatory genes plus the elements they regulate, perform specific functions
- Switches permit or forbid the activity of whole sub-circuits
- Gene batteries are groups of genes required for particular cell functions; they are controlled by a small set of transcriptional drivers
- Segments of DNA a few hundred base pairs long, called cis-regulatory elements, control expression of the genes near them
- Signals are deployed between one cell and another using cis-regulatory elements
- Erwin, Douglas H., Eric H. Davidson. February 2009. The evolution of hierarchical gene regulatory networks. Nature Reviews Genetics, Vol. 10, pp. 141-148.
Mutation-natural selection could no more build the vast, intricate networks in living creatures than a beaver could build the Hoover dam.

"Before Darwin came along, it seemed absurd to almost everyone that the beauty and complexity of the living world could have come into being without a designer." (p. 250)
"...the complexity, beauty, and 'purposefulness' of living things must have seemed too obviously designed by an intelligent creator. So it requires a major leap of courage to consider anything else I mean intellectual courage to contemplate the apparently ridiculous." (p. 267) - From Dawkins' 2019 book "Outgrowing God"
Inventing machines without a designer is ridiculous. All machines have been engineered. One of millions of design examples is how tuna use hydraulics to maneuver, and "employ a hydraulic system that is similar to an engineers design."
"Highly streamlined bodies (like those of tuna) without fins are unstable, as the aerodynamic center is forward of the head, often by a full body length. This necessitates the use of hydrofoils in the rear part of the body to bring the aerodynamic center close to its center of mass."
"By changing the sweep angle of the front dorsal, second dorsal, and anal fins, the tuna can rapidly move its aerodynamic center, either forward to reduce stability and hence increase maneuverability, or backward to increase stability." "Reducing the sweep angle increases the effective aspect ratio and the lift per unit angle of attack."
|
|
|
"The lymphatic circulatory system in tuna serves as a hydraulic system that actively changes the sweep angle of their second dorsal (back) and anal (underside) fins." "Such changes are used consistently during maneuvering and transient motion, especially for sharp turns." |
Although "the lymphatic system has a key role in other biological processes (primarily in the immune system)", "the muscular system alone cannot provide and sustain with accuracy the forces required to raise the fin rays and thereby change the sweep angle of the fins. Instead, the inclinator muscles, located around the base of the fins, contract to squeeze compressible vascular channels, pushing lymph through the incompressible vascular sinus to fill expandable vascular channels between the fin rays."
"Continuous adjustment of the sweep angle is found in other swimming animals that employ lift-based propulsion, so this could be a mechanism of wider applicability."
"The shape-changing fins of fish are of great interest to engineers developing the locomotion and maneuvering capabilities of underwater and aerial systems."
- Triantafyllou, Michael S. 21 July 2017. Tuna fin hydraulics inspire aquatic robotics - How tuna control fin shape while making sharp turns suggests optimum engineering design. Science, Vol. 357, No. 6348 DOI:10.1126/science.aan8354
To the next level
We are now light-years beyond the simplistic notions of Darwinism; even systems biology is overwhelmed.
"Increasingly, leaders in the fields of systems biology realise that it is our inability to understand and model the true underlying complexity of whole biological systems (as opposed to their reduced parts) that is holding back deep physiological understanding of organisms: truly understanding biology involves getting to grips with unimaginable complexity". - Moore, Andrew. 2012. Bringing systems biology to the clinic: An acute case. Bioessays Vol. 35, No. 1, pg. 1.
"The protein p53, for example, was discovered in 1979." "It soon gained notoriety as a tumor suppressor - a 'guardian of the genome' that stifles cancer growth by condemning genetically damaged cells to death. Few proteins have been studied more than p53."
"Researchers now know that p53 binds to thousands of sites in DNA, and some of these sites are thousands of base pairs away from any genes. It influences cell growth, death, structure and DNA repair. It also binds to numerous other proteins." "Through a process known as alternative splicing, p53 can take nine different forms, each of which has its own activities and chemical modifiers. Biologists are now realizing that p53 is also involved in processes beyond cancer, such as fertility and very early embryonic development."
Research "has dismantled old ideas about signaling 'pathways', in which proteins such as p53 would trigger a defined set of downstream consequences. 'When we started out, the idea was that signaling pathways were fairly simple and linear,' says Tony Pawson, a cell biologist at the University of Toronto in Ontario. 'Now, we appreciate that the signaling information in cells is organized through networks of information rather than simple discrete pathways. It's infinitely more complex.' "
"Systems biology was supposed to help scientists make sense of the complexity. The hope was that by cataloguing all the interactions in the p53 network, or in a cell, or between a group of cells, then plugging them into a computational model, biologists would glean insights about how biological systems behaved."
Unfortunately, "there is no way to gather all the relevant data about each interaction". "In many cases, the models themselves quickly become so complex that they are unlikely to reveal insights about the system, degenerating instead into mazes of interactions". "Many of the mechanisms and principles governing inter- and intracellular behavior are still a mystery." - Hayden, Erika Check. 1 April 2010. Life is Complicated. Nature, Vol. 464, pp. 664-667.
When cells repair damaged DNA using so-called "replicate DNA" (for making copies) or "transcribe DNA" (the first step in making a protein), the parts are rapidly assembled from a pool of parts floating in the nucleus of a cell to form "factories". The size of a repair center is according to the amount of damage it has to repair. "A replication factory persists for a few minutes before it disassembles. A new factory is then assembled... immediately adjacent to the previous one". Whether it is repair, replication, or transcription, once the job is done the "factories" disassemble and the parts float back into the pool.32
Chromatin is DNA packaged into chromosomes. Chromatin is in constant motion.3 Different sections along DNA are apparently guided to each other directly and rapidly, forming loops.11 Chromatin loops are very common. Loops range in size from thousands to hundreds-of-thousands of bases long. Loops bring together genes and their regulators to form "transcription hubs" where transcription can occur.32 "Long-range interactions can occur over very large genomic distances, up to tens of megabases". "Interactions occur not only along chromosomes, but also between them." "Chromosomes extensively interact with each other"42, and neighboring chromosomes intermingle.32
There is a "bewildering complexity in long-range communication among a variety of genomic elements across chromosomes and the genome."42
The Bottom Line
Evolutionists
assume evolution is true, then write endlessly about when
and where it happened, rates and lineages, etc. But
if macroevolution is physically impossible in the real
world, and it is, then all the rest is fantasy. There are only two possibilities. Either every
part of every
living thing arose by random chance, or an intelligence designed them.
|
|
|
It is now clear that the theory of evolution's only mechanism for building new parts and creatures, mutation-natural selection, is totally, utterly, pathetically inadequate. |
In spite of overwhelming evidence that the theory of evolution is dead wrong, many are not ready to throw in the towel. They desperately hope that some natural process will be found that causes things to fall together into organized complexity. These are people of great faith. And they are so afraid of connecting God with science that, like the Japanese Army of World War II, they would rather die than surrender. Unfortunately, the staunchest defenders sit in places of esteem and authority as professors, scientists, and editors, and have the full faith of the news media. The public is naturally in awe of their prestige. But once the facts are understood it becomes obvious that the theory of evolution is long overdue for the trash can, and to perpetuate it is fraud. Perhaps it made sense for what was known when On the Origin of Species was published in 1859, but not today.
Many scientists are with us
The only tactic left to evolutionists
is to ridicule their critics as simpletons who don't understand how their pet
theory really works. Here is a link to a roster of
hundreds of professionals whose advanced academic degrees certify that they thoroughly
understand evolution theory. They also have the
courage to defy the high priests of academia by voluntarily adding
their names to a skeptics
list against Darwinism.
Richard Dawkins
- obsolete
Nathaniel
Comfort is professor of the history of medicine at Johns Hopkins University in
Baltimore, Maryland. He reviewed Richard Dawkins' autobiography Brief
Candle in the Dark: My Life in Science (2015) for the evolutionist
journal Nature, saying in part:
"Much of Dawkinss research has been... writing programs for evolutionary simulations. In his simulations, life is utterly determined by genes, which specify developmental rules and fixed traits such as colour. The more lifelike his digital animals ('biomorphs') become, the more persuaded he is that real genes work in roughly the same way."
"A curious stasis underlies Dawkinss thought. His biomorphs are grounded in 1970s assumptions. Back then, with rare exceptions, each gene specified a protein and each protein was specified by a gene. The genome was a linear text - a parts list or computer program for making an organism - insulated from the environment, with the coding regions interspersed with 'junk'."
"Todays genome is much more than a script: it is a dynamic, three-dimensional structure, highly responsive to its environment and almost fractally modular. Genes may be fragmentary, with far-flung chunks of DNA sequence mixed and matched in bewildering combinatorial arrays. A universe of regulatory and modulatory elements hides in the erstwhile junk."
"Dawkinss synopsis shows that he has not adapted to this view." "The microbiome and the 3D genome go unnoticed. Epigenetics is an 'interesting, if rather rare, phenomenon' enjoying its 'fifteen minutes of pop science voguery' ". "Dawkins adheres to a deterministic language of 'genes for' traits."
"In the early 2000s, he saltated from popularizer into evangelist." Along with Christopher Hitchens, Daniel Dennett and Sam Harris, his writings "form the scripture of the 'new atheism', a fundamentalist sect that has mounted a scientistic crusade against all religion.
"For a time, Dawkins was a rebellious scientific rock star. Now, his critique of religion seems cranky, and his immovably genocentric universe is parochial." - Comfort, Nathaniel. 10 September 2015. Dawkins, redux. Nature, Vol. 525, No. 7568, pp. 184-185.
Some revealing quotes
|
|
Philip S. Skell, a member of the National Academy of Sciences, wrote in the August 29, 2005 edition of The Scientist: "I recently asked more than seventy eminent researchers if they would have done their work differently if they had thought Darwin's theory was wrong. The responses were all the same: No. I also examined the outstanding discoveries of the past century: the discovery of the double helix; the characterization of the ribosome; the mapping of genomes; research on medications and drug reactions; improvements in food production and sanitation; the development of new surgeries; and others. I even queried biologists working in areas where one would expect the Darwinian paradigm to have to have most benefited research, such as the emergence of resistance to antibiotics and pesticides. Here, as elsewhere, I found that Darwin's theory had provided no discernible guidance, but was brought in, after the breakthroughs, as an interesting narrative gloss." - Philip S. Skell. August 29, 2005. Why Do We Invoke Darwin? The Scientist, Vol. 19, No. 16, p. 10. |
Philip S. Skell was Evan Pugh Professor of Chemistry,
Emeritus at Penn State University. He is sometimes called "the
father of carbene chemistry" in organic chemistry, and is widely
known for the "Skell Rule", which was first applied to carbenes
- the "fleeting species" of carbon. The rule, which
predicts the most probable pathway through which certain chemical
compounds will be formed, found use throughout the pharmaceutical and
chemical industries. He said that during World War II "I was
personally associated with an antibiotics research group, engaged in the
full range of activities, from finding organisms which inhibited bacterial
growth to the isolation and proof of structure of the antibiotics they
produced." Professor Skell died Nov. 21, 2010.
|
|
Ernst Chain (1906-1979) and two others were awarded the 1945 Nobel Prize for Physiology or Medicine. Chain identified the structure of penicillin, and isolated the active substance. He is considered to be one of the founders of the field of antibiotics. Concerning Darwin's theory of evolution, Chain found it to be "a very feeble attempt" to explain the origin of species based on assumptions so flimsy that "it can hardly be called a theory."A |
He saw the reliance on chance mutations as a
"hypothesis based on no evidence and irreconcilable with the
facts."B He wrote: "These classic evolutionary
theories are a gross oversimplification of an immensely complex and intricate
mass of facts, and it amazes me that they were swallowed so uncritically and
readily, and for such a long time, by so many scientists without a murmur of
protest."B Chain concluded that he "would
rather believe in fairies than in such wild speculation" as Darwinism.A He was born in Berlin, Germany, and obtained his
Ph.D. in biochemistry and physiology there. He worked as a research
scientist at Cambridge (also studying for a Ph.D. there), at Oxford University
until 1948, and then as a professor and researcher at several other
universities. In 1938, Chain came across Alexander Fleming's 1929 paper
on penicillin, and showed it to his colleague Howard Florey. In their
research, Chain isolated and purified penicillin. - Jerry
Bergman, Ph.D. April 2008. Ernst Chain: Antibiotics Pioneer. Acts&Facts,
Vol. 37, No. 4, pp. 10-12.
A. Clark, R.W. 1985. The Life of Ernst Chain: Penicillin and Beyond. New
York: St. Martin's Press, p. 147.
B. Chain, E. 1970. Social Responsibility and the Scientist in Modern
Western Society (Robert Waley Cohen memorial lecture). London: The Council of
Christians and Jews, p. 25.
|
|
Perry Marshall has a degree in Electrical Engineering. Hes consulted in over 300 industries, from computer hardware and software to biotech and health care, and is the author of the book Evolution 2.0. This is from his website http://cosmicfingerprints.com/dna-atheists/ "For well over 5 years, I successfully advanced the Information Theory argument for design in DNA on Infidels, which from 2005-2010 was the world's largest Atheist discussion forum." [It has since disappeared.] |
"In over five years, the Infidels failed to refute the information theory argument for design in biology."
"Gentlemen: The starting point of this discussion is my central thesis, which is: 1) DNA is not merely a molecule with a pattern; it is a code, a language, and an information storage mechanism. 2) All codes are created by a conscious mind; there is no natural process known to science that creates coded information. 3) Therefore DNA was designed by a mind. If you can provide an empirical example of a code or language that occurs naturally, youve toppled my proof. All you need is one. - Perry Marshall"
"In the course of a very detailed and vigorous discussion my argument did not suffer the slightest injury. There were six major counter-arguments to information as proof of intelligent design.
1. The objection that DNA is not a code (it is, by universal definition)
2. The objection that information is not real (it is, because it produces real effects)
3. The objection that information has no objective meaning (it does, because a message produces results that are just as objective and specific as the message itself)
4. The objection that random processes can create information (they cant)
5. The objection that codes do occur naturally (they dont)
6. The objection that the nature of the Designer cannot be determined (in very broad terms, it can)"
"The sequence of base pairs in DNA is a code. All codes that we know the origin of come from a mind. Therefore DNA came from a mind."
"The objection to this statement has been that the conclusion is reached inductively. Complaints have been lodged that inductive reasoning is inherently unreliable. But we do observe that the laws of thermodynamics and in fact the majority of known scientific laws are determined inductively and not deductively."
"Lets not forget that the entire enterprise of scientific inquiry during the last 500 years has been the ongoing discovery of underlying order, not the assumption of accident."
"Thus we have, right here on the Infidels discussion forum, after more than 300 posts, robust evidence that life was intelligently designed."
"It is not possible for me to persuade people to believe in God if they do not want to; that is not my job. But one can hope that some will follow the evidence, wherever it leads."
|
|
"My experiences with science led me to God. To be forced to believe only one conclusion -- that everything in the universe happened by chance -- would violate the very objectivity of science itself. Certainly there are those who argue that the universe evolved out of a random process, but what random process could produce the brain of a man or the system of the human eye? Some people say that science has been unable to prove the existence of a Designer... They challenge science to prove the existence of God. But, must we really light a candle to see the sun?" --Wernher von Braun, 1912 1977 |
- "Wernher von Braun is, without doubt, the greatest rocket scientist in history. His crowning achievement, as head of NASA's Marshall Space Flight Center, was to lead the development of the Saturn V booster rocket that helped land the first men on the Moon in July 1969." --From NASA's webpage: http://earthobservatory.nasa.gov/Features/vonBraun/
Richard C. Strohman, professor emeritus of molecular and cell biology at Berkeley, and an evolutionist, wrote in the March 1997 edition of Nature Biotechnology: "There is a striking lack of correspondence between genetic and evolutionary change. Neo-Darwinian theory predicts a steady, slow continuous, accumulation of mutations (microevolution) that produces a progressive change in morphology leading to new species, genera, and so on (macroevolution). But macroevolution now appears to be full of discontinuities (punctuated evolution), so we have a mismatch of some importance. That is, the fossil record shows mostly stasis, or lack of change, in a species for many millions of years; there is no evidence there for gradual change even though, in theory, there must be a gradual accumulation of mutations at the micro level." "We currently have no adequate explanation for stasis or for punctuated equilibrium in evolution, or for higher order regulation in cells." "We seem to lack any scientific basis with which to explain, for example, evolution." "Not necessarily so. It does suggest, however, that our evolutionary theory is incomplete." "The theory is in trouble because it insists on locating the driving force solely in random mutations." "It is becoming clear that sequence information in DNA, by itself, contains insufficient information for determining how gene products (proteins) interact to produce a mechanism of any kind. The reason is that the multicomponent complexes constructed from many proteins are themselves machines with rules of their own; rules not written in DNA." "The rules... of brain formation are not reducible to genetic maps and to the rules of genetic theory. Each higher level of organization has its own rules, and there is no continuous gradual transition from one level or hierarchy to the other." "We have been lulled into reasoning that if the gene theory works at one level--from DNA to protein--it must work at all higher levels as well. We have thus extended the theory of the gene to the realm of gene management. But gene management is an entirely different process, involving interactive cellular processes that display a complexity that may only be described as transcalculational, a mathematical term for mind boggling." "Understanding of complex function may in fact be impossible without recourse to influences outside of the genome." - Richard C. Strohman. March 1997. The coming Kuhnian revolution in biology. Nature Biotechnology, Vol. 15, pp. 194-200.
Sean B. Carroll, of the Medical Institute and Laboratory of Molecular Biology at the University of Wisconsin--Madison, wrote in a 2001 edition of Nature: "A long-standing issue in evolutionary biology is whether the processes observable in extant populations and species (microevolution) are sufficient to account for the larger-scale changes evident over longer periods of life's history (macroevolution). Outsiders to this rich literature may be surprised that there is no consensus on this issue." - Sean B. Carroll. 8 February 2001. Nature, Vol. 409, p. 669.
A symposium on evolution was held at the European Molecular Biology Laboratory in Heidelberg, Germany in November 2001, organized by PhD students. The meeting report says that "the symposium ended with a panel discussion about questions of microevolution (evolution within the species) and macroevolution (evolution after speciation). The issue at stake was whether extrapolation from the selection theory operating on organisms is sufficient to explain all patterns of macroevolution. In other words, do we need an independent body of theory to explain the changes occurring above, as opposed to at, the species level? There was no general agreement among the panel members. It seems that the jury is still out on this important question." - Gaspar Jekely. 2002. Meeting report - Evolution in a nutshell. European Molecular Biology Organization reports, Vol. 3, No. 4, pp. 307-311.
"Biology has been re-integrated
twice already, first by Darwin in 1859 and then during the 'Modern
Synthesis' of the 1920s and 1930s. In both cases, the success
of these syntheses rested in part on ignorance. Charles Darwin
could reasonably integrate biology in the 19th
Century on a relatively elegant evolutionary foundation partly because
a great deal was not yet known about cellular and biochemical machinery."
"Like Darwin's synthesis, the form of the Modern Synthesis
was shaped in part by ignorance of important features of life that
were at the time unknown to science. Specifically, the molecular
biology of the cell remained largely unknown." "The
view of life that most biologists had from 1935 to 1965 was highly
simplified. Some of the assumptions at the foundation of the
Modern Synthesis started to crumble in the 1970s. Common mid-20th
Century assumptions about how cells, organisms, and species work
have thus been undermined." "This might seem like
reason for despair about the future of biology, but there are two
mitigations to consider. First, this complexity was always
there. Darwin and many later biologists realized that their
simple models were erected like piers over swampy ground. They
just didn't know how deep the muck was. Second, we now have
powerful genomic tools for addressing complex phenomena throughout
biology." "Some may feel that the view of life supplied
by nascent 21st
Century biology is painfully complicated, if not perverse. For
our part, we think that the historical complexity and versatility
that we now know to characterize life are inspiring and challenging."
![]() "The fundamental landscape of biology is undergoing a
major upheaval, much as it did in the first decades of the 20th
Century. This upheaval will take time to fully reveal its
implications." - Michael R. Rose, Todd H. Oakley. 24
November 2007. The new biology: beyond the Modern Synthesis. Biology
Direct, 2:30, 17 pages (published online). Michael Rose is an evolutionary biologist
at the University of California, Irvine.
"The fundamental landscape of biology is undergoing a
major upheaval, much as it did in the first decades of the 20th
Century. This upheaval will take time to fully reveal its
implications." - Michael R. Rose, Todd H. Oakley. 24
November 2007. The new biology: beyond the Modern Synthesis. Biology
Direct, 2:30, 17 pages (published online). Michael Rose is an evolutionary biologist
at the University of California, Irvine.
"The philosophers of Greece have made much ado to explain nature... Those who were too ignorant to rise to a knowledge of a God could not allow that an intelligent cause presided at the birth of the universe... Some had recourse to material principles and attributed the origin of the universe to the elements of the world. Deceived by their inherent atheism, it appeared to them that nothing governed or ruled the universe, and that all was given up to chance." - 370 AD Saint Basil the Great, Bishop of Caesarea Mazaca in Cappadocia (Turkey), Homily I on the Hexaemeron.
|
|
|
"The harmony of natural law reveals an Intelligence of such superiority that, compared with it, all the systematic thinking and acting of human beings is an utterly insignificant reflection." - Albert Einstein. 1931. Ideas and Opinions - The World As I See It. New York, Bonanza Books. Page 40. |
"Radiation" means new creatures evolved. "The hypothesis that destructive mass extinctions enable creative evolutionary radiations (creative destruction) is central to classic concepts of macroevolution." However
"Among the 5% most significant periods of disruption, we identify the 'big five' mass extinction events, seven additional mass extinctions, two combined mass extinctionradiation events and 15 mass radiations. In contrast to narratives that emphasize post-extinction radiations, we find that the proportionally most comparable mass radiations and extinctions (such as the Cambrian explosion and the end-Permian mass extinction) are typically decoupled in time, refuting any direct causal relationship between them."
"This analysis shows that the most comparable mass radiations and extinctions are in general temporally decoupled, strongly arguing against an immediate causal connection between them. In particular, the proportionately most extreme mass extinctions were, necessarily, not accompanied by a radiation of comparable scope within the same 1 million year time window. Nor are the mass extinctions generally observed to be closely followed by a mirroring mass radiation, which would be predicted by hypotheses of vacation of niches and direct replacement, for example. Instead, the events in Phanerozoic history that have created proportionately the most diversity (including mass radiations at the beginning of the Cambrian, Carboniferous, Late Ordovician and early Cretaceous) have generally occurred at times that were widely separated from the mass extinction events. One notable exception to this temporal decoupling of mass extinction and radiation is the end-Permian mass extinction at 252 Ma, which was followed closely by two significant radiation events at 251 and 247 Ma." - Goedel, Alexander, Fredrik Lanner. 17 November 2021. A peek into the black box of human embryology. Nature, Vol. 600, No. 7888, pp. 223-224, DOI:10.1038/d41586-021-03381-x
"Origin
of Life" research, continued
The theory
of evolution says life started from raw chemicals. Evolutionists
long ago handed the problem off to specialists, trusting that they
would come up with something. The specialists have spent many
frustrating decades trying to figure out how DNA assembled itself.
They have two approaches to the problem, and those on one
side think the other side is wrong. Here is the essence of
both views, synthesized from two research papers:
The mainstream prebiotic evolutionary scenario is the "RNA world".S "Textbooks often assert that life began with specialized complex molecules, such as RNA, that are capable of making their own copies. This scenario has serious difficulties, but an alternative has remained elusive."S "We do not know how the transition to digitally encoded information has happened in the originally inanimate world; that is, we do not know where the RNA world might have come from."V
"An alternative appears to be necessary for the RNA-centric paradigm of the origin of life."S "No known cellular constituent is capable of self-replication in pure form. Even DNA is absolutely dependent on other cellular components for making its own copies."S "One is compelled to consider an alternative: that self-replication has never been a property of individual molecules, but rather one of molecular ensembles."S"The crucial origin of life question then becomes how natural selection was initiated by some molecular assortments, irrespective of their exact chemistry."S "Life on our planet could have begun as a random chemistry melting pot, a 'garbage-bag world' with myriads of different chemical configurations."S "A complex chain of evolutionary events, yet to be deciphered, could then have led to the common ancestors of today's free-living cells, and to the appearance of DNA, RNA and protein enzymes."S
"Was a network of chemical reactions able to increase in complexity and eventually undergo Darwinian selection?"V "We demonstrate here that replication of compositional information is so inaccurate that fitter compositional genomes cannot be maintained by selection and, therefore, the system lacks evolvability."V "The computed population dynamics of growing noncovalent molecular assemblies that undergo splitting when a critical size is reached clearly illustrates that compositional assemblies do not evolve."V "We conclude that this fundamental limitation of ensemble replicators cautions against metabolism-first theories of the origin of life."V "We now feel compelled to abandon compositional inheritance as a jumping board toward real units of evolution."VS -- Segre, Daniel, Doron Lancet. 2000. Composing life. European Molecular Biology Organization (EMBO) Reports, Vol. 1, No. 3, pp. 217-222.
V -- Vasas, Vera, Eors Szathmary, Mauro Santos. January 26, 2010. Lack of evolvability in self-sustaining autocatalytic networks constraints metabolism-first scenarios for the origin of life. Proceedings of the National Academy of Sciences of the United States of America (PNAS), Vol. 107, No. 4, pp. 1470-1475.
So neither approach works.
Here are excerpts from candid reports by two scientists who have spent many years in "origin of life" research. These men support evolution, but insist that experimental evidence back up every claim.
This is "what has been called the NASA definition of life: Life is a self-sustained chemical system capable of undergoing Darwinian evolution." "Richard Dawkins elaborated on this image of the earliest living entity in his book The Selfish Gene: 'At some point a particularly remarkable molecule was formed by accident. We will call it the Replicator. It may not have been the biggest or the most complex molecule around, but it had the extraordinary property of being able to create copies of itself.' When Dawkins wrote these words 30 years ago, DNA was the most likely candidate for this role." "Unfortunately... DNA replication cannot proceed without the assistance of a number of proteins". So "which came first, the chicken or the egg? DNA holds the recipe for protein construction. Yet that information cannot be retrieved or copied without the assistance of proteins. Which large molecule, then, appeared first in getting life started--proteins (the chicken) or DNA (the egg).?"
"A possible solution appeared when attention shifted to a new champion--RNA." According to this view, "life began with the appearance of the first RNA molecule. In a... 1986 article, Nobel Laureate Walter Gilbert of Harvard University wrote in the journal Nature: 'One can contemplate an RNA world, containing only RNA molecules that serve to catalyze the synthesis of themselves. The first step of evolution proceeds then by RNA molecules performing the catalytic activities necessary to assemble themselves from a nucleotide soup.' In this vision, the first self-replicating RNA that emerged from non-living matter carried out the functions now executed by RNA, DNA and proteins." "Perhaps two-thirds of scientists publishing in the origin-of-life field... still support the idea that life began with the spontaneous formation of RNA or a related self-copying molecule."
"How did that first self-replicating RNA arise?" Most people know of an "experiment published in 1953 by Stanley Miller. He applied a spark discharge to a mixture of simple gases that were then thought to represent the atmosphere of the early Earth. Two amino acids of the set of 20 used to construct proteins were formed in significant quantities, with others from that set present in small amounts." "Some writers have presumed that all of life's building blocks could be formed with ease in Miller-type experiments and were present in meteorites and other extraterrestrial bodies. This is not the case."
"A careful examination of the results of the analysis of several meteorites led the scientists who conducted the work to a different conclusion: inanimate nature has a bias toward the formation of molecules made of fewer rather than greater numbers of carbon atoms, and thus show no partiality in favor of creating the building blocks of our kind of life." "RNA's building blocks, nucleotides, are complex substances as organic molecules go." "Amino acids, such as those produced or found in these experiments, are far less complex than nucleotides". "No nucleotides of any kind have been reported as products of spark discharge experiments or in studies of meteorites."
"To rescue the RNA-first concept from this otherwise lethal defect, its advocates have created a discipline called prebiotic synthesis. They have attempted to show that RNA and its components can be prepared in their laboratories in a sequence of carefully controlled reactions." Finding "a specific organic chemical in any quantity... would justify its classification as 'prebiotic,' a substance that supposedly had been proved to be present on the early Earth. Once awarded this distinction, the chemical could then be used in pure form, in any quantity, in another prebiotic reaction. The products of such a reaction would also be considered 'prebiotic' and employed in the next step in the sequence." "Unfortunately, neither chemists nor laboratories were present on the early Earth to produce RNA." "The analogy that comes to mind is that of a golfer, who having played a golf ball through an 18-hole course, then assumed that the ball could also play itself around the course in his absence. He had demonstrated the possibility of the event; it was only necessary to presume that some combination of natural forces (earthquakes, winds, tornadoes and floods, for example) could produce the same result, given enough time."
"Many chemists, confronted with these difficulties, have fled the RNA-first hypothesis as if it were a building on fire. One group, however, still captured by the vision of the self-copying molecule, has opted for an exit that leads to similar hazards. In these revised theories, a simpler replicator arose first and governed life in a 'pre-RNA world.' Variations have been proposed in which the bases, the sugar or the entire backbone of RNA have been replaced by simpler substances, more accessible to prebiotic syntheses. Presumably, this first replicator would also have the catalytic capabilities of RNA. Because no trace of this hypothetical primal replicator and catalyst has been recognized so far in modern biology, RNA must have completely taken over all of its functions at some point following its emergence."
"Further, the spontaneous appearance of any such replicator without the assistance of a chemist faces implausibilities that dwarf those involved in the preparation of a mere nucleotide soup. Let us presume that a soup enriched in the building blocks of all of these proposed replicators has somehow been assembled, under conditions that favor their connection into chains. They would be accompanied by hordes of defective building blocks, the inclusion of which would ruin the ability of the chain to act as a replicator." "There is no reason to presume that an indifferent nature would not combine units at random".
"Probability calculations could be made, but I prefer a variation on a much-used analogy. Picture a gorilla (very long arms are needed) at an immense keyboard connected to a word processor. The keyboard contains not only the symbols used in English and European languages but also a huge excess drawn from every other known language and all of the symbol sets stored in a typical computer. The chances for the spontaneous assembly of a replicator in the pool I described above can be compared to those of the gorilla composing, in English, a coherent recipe for the preparation of chili con carne. With similar considerations in mind, Gerald F. Joyce of the Scripps Research Institute and Leslie Orgel of the Salk Institute concluded that the spontaneous appearance of RNA chains on the lifeless Earth 'would have been a near miracle.' I would extend this conclusion to all of the proposed RNA substitutes that I mentioned above." "Nobel Laureate Christian de Duve has called for 'a rejection of improbabilities so incommensurably high that they can only be called miracles, phenomena that fall outside the scope of scientific inquiry.' DNA, RNA, proteins and other elaborate large molecules must then be set aside as participants in the origin of life."
What is left? Theories that "employ a thermodynamic rather than a genetic definition of life, under a scheme put forth by Carl Sagan in the Encyclopedia Britannica: A localized region which increases in order (decreases in entropy) through cycles driven by an energy flow would be considered alive." "I estimate that about a third of the chemists involved in the study of the origin of life subscribe to theories based on this idea."
It requires: "1) A boundary... to separate life from non-life." "2) An energy source". "3) A coupling mechanism must link the release of energy to the organization process that produces and sustains life. The release of energy does not necessarily produce a useful result. Chemical energy is released when gasoline is burned within the cylinders of my automobile, but the vehicle will not move unless that energy is used to turn the wheels. A mechanical connection, or coupling, is required." "4) A chemical network must be formed, to permit adaptation and evolution" "on a path that leads to increased organization." "5) The network must grow and reproduce." "We can imagine, on the early Earth, a situation where many startups of this type occur, involving many alternative driver reactions and external energy sources. Finally, a particularly hardy one would take root and sustain itself." "A system of reproduction must eventually develop." "Once independent units were established, they could evolve in different ways and compete with one another for raw materials; we would have made the transition from life that emerges from nonliving matter through the action of an available energy source to life that adapts to its environment by Darwinian evolution." "Many further steps in evolution would be needed to 'invent' the elaborate mechanisms for replication and specific protein synthesis that we observe in life today." They "would not reveal the specific events that led to the familiar DNA-RNA-protein-based organisms of today."
"Systems
of the type I have described usually have been classified under
the heading 'metabolism first', which implies that they do not contain
a mechanism for heredity. In other words, they contain no
obvious molecule or structure that allows the information stored
in them (their heredity) to be duplicated and passed on to their
descendants." "Over the years, many theoretical
papers have advanced particular metabolism first schemes, but relatively
little experimental work has been presented in support of them."
"They have not yet demonstrated the operation of a complete
cycle or its ability to sustain itself and undergo further evolution.
A 'smoking gun' experiment displaying those three features
is needed to establish the validity of the small molecule approach."
Shapiro, Robert. June 2007. A Simpler Origin
for Life. Scientific American, Vol. 296, pp. 24-31.
Robert Shapiro,
Ph.D. Harvard, is professor emeritus of chemistry and senior research
scientist at New York University. He is author or co-author
of over 125 publications, primarily in the area of DNA chemistry.
In 2004 he was awarded the Trotter Prize in Information, Complexity
and Inference. Shapiro has been involved in the search for
origin of life mechanisms, and has written four books on the subject
for the general public.
"The feasibility of any particular proposed prebiotic cycle must depend on arguments about chemical plausibility, rather than on a decision about logical possibility." The metabolic cycles that have been identified by biochemists are of two kinds: simple cycles and autocatalytic cycles. The citric acid cycle" is an example of a simple cycle. "The reverse citric acid cycle" is an example of an autocatalytic cycle. "Each molecule of citric acid introduced into the cycle results... in the generation of two molecules of citric acid." "That is why the cycle is described as autocatalytic." "The proposal that the reverse citric acid cycle operated... on the primitive Earth has been a prominent feature of some scenarios for the origin of life."
"A different kind of autocatalytic cycle, which has no analog in biochemistry, has been hypothesized by Stuart Kauffman to self-organize spontaneously whenever amino acids condense together to form peptides." "Could prebiotic molecules and catalysts plausibly have the attributes... to make the self-organization of the cycles possible?"
"The identification of a cycle of plausible prebiotic reactions is a necessary but not a sufficient step toward the formulation of a plausible self-organizing prebiotic cycle. The next, and more difficult step, is justifying the exclusion of side reactions that would disrupt the cycle." "It is not completely impossible that sufficiently specific mineral catalysts exist for each of the reactions of the reverse citric acid cycle, but the chance of a full set of such catalysts occurring at a single locality on the primitive Earth in the absence of catalysts for disruptive side reactions seems remote in the extreme."
"It has sometimes been implied or claimed that [autocatalytic] cycles are not only stable, but also are capable of evolving to form nonenzymatic networks of great complexity. Genetic materials are then seen as late additions to already fairly complex evolved life forms. According to this view, a genetic material merely adds stability to systems that already have a substantial 'information content'. "
"One way of achieving something useful might be to use one of the constituents of the core cycle as the starting point of a second independent autocatalytic cycle." "Another suggestion that might be explored is the possibility of a side reaction generating a catalyst for one of the reactions of the core cycle." "However, neither of these possibilities, nor any others with which I am familiar explains how a complex, interconnected family of cycles capable of evolution could arise or why it should be stable." "What is essential, therefore, is a reasonably detailed description, hopefully supported by experimental evidence, of how an evolvable family of cycles might operate." "Without such a description, acceptance of the possibility of complex nonenzymatic cyclic organizations that are capable of evolution can only be based on faith, a notoriously dangerous route to scientific progress."
"Kauffman takes it for granted that if it is possible to write down on paper a closed peptide cycle and a set of catalyzed ligations leading from monomeric amino acids to the peptides of the cycle, then that cycle would self-organize spontaneously and come to dominate the chemistry of a reaction system. This... is unlikely because peptide molecules do not have the properties that Kauffman assigns to them." "I have also explored a number of alternative systems with different numbers of amino acids or with inputs of random families of short peptides, and I find that they all encounter similar or more severe difficulties."
"Kauffman assumes that, in sufficiently concentrated solution, the naturally occurring amino acids or some subset of them would condense spontaneously to form a mixture of long peptides in substantial yield. In practice, this would not happen." "The catalytic properties of enzymes are remarkable. They not only accelerate reaction rates by many orders of magnitude, but they also discriminate between potential substrates that differ very slightly in structure. Would one expect similar discrimination in the catalytic potential of peptides of length ten or less? The answer is clearly 'no', and it is this conclusion that ultimately undermines the peptide cycle theory."
"Protein catalysis is dependent on the stable three-dimensional structures of enzymes and enzyme-substrate complexes. Highly specific catalytic activity could only be expected from short peptides if they, too, adopted stable structures." "In fact, short peptides rarely form stable structures, and when they do, the structures are only marginally stable. The synthesis of a decapeptide that would catalyze the ligation in the correct order of two particular pentapeptides out of a mixture of ten pentapeptides that are required to form the five cycle components, while failing to bring about any of the other possible ligations, would present an extremely difficult challenge to peptide chemistry. It seems certain that the additional requirement that this peptide should also catalyze specifically many of the reactions leading from amino acids to the pentamer precursors of the decamers of the cycle could never be met. Of course, the decamers need not be formed only from pairs of pentamers, but the difficulties are no less severe for more complex synthetic networks. There are a number of possible ways in which this difficulty might be circumvented, but none seems relevant to the origin of life." "It is unlikely, therefore, that Kauffman's theory describes any system relevant to the origin of life."
"It is essential to subject metabolist proposals to the same kind of detailed examination and criticism that has rightly been applied to genetic theories." "Because little experimental work has been attempted, appraisal must be based on chemical plausibility." "The lack of a supporting background in chemistry is even more evident in proposals that metabolic cycles can evolve to 'life-like' complexity. The most serious challenge to proponents of metabolic cycle theories--the problems presented by the lack of specificity of most nonenzymatic catalysts--has, in general, not been appreciated. If it has, it has been ignored."
"Theories of the origin of life based on metabolic cycles cannot be justified by the inadequacy of competing theories: they must stand on their own." "Experimental proof that such cycles are stable against the challenge of side reactions is even more important." "The prebiotic syntheses that have been investigated experimentally almost always lead to the formation of complex mixtures. Proposed polymer replication schemes are unlikely to succeed except with reasonably pure input monomers. No solution of the origin-of-life problem will be possible until the gap between the two kinds of chemistry is closed." "Solutions offered by supporters of geneticist or metabolist scenarios that are dependent on 'if pigs could fly' hypothetical chemistry are unlikely to help." - Orgel, Leslie E. January 2008. The Implausibility of Metabolic Cycles on the Prebiotic Earth. Public Library of Science (PLoS) Biology, Vol. 6, No. 1, e18, pp. 5-13.
Leslie E. Orgel, Ph.D. Oxford, was a biochemist who studied life on primitive Earth. He conducted research at Cambridge, the University of Chicago, the California Institute of Technology, and later joined the Chemical Evolution Laboratory of the Salk Institute for Biological Studies in San Diego, California. He died at age 80 in October 2007. The above article was published posthumously.
After the
"tree of life"
In a paper about bacteria,
two evolutionary biologists write, "we cannot rely exclusively
on traditional genealogical relationships." "A single
taxonomy will be likely to provide an overly coarse picture".
It should be replaced by "more taxonomies based on real
biological processes". "Discarding all but one of
these process-based taxonomies would be comparable to reducing a
person's identity to a single aspect of his or her life, even though
he or she might have an effective role in many organizations: professional,
artistic, sports, family and so on. To avoid overlooking any
of the natural groups, it seems legitimate to propose - rather than
a single taxonomy of microbial species - many taxonomies".
"We suggest giving up the unique hierarchy as the reference
classification system and instead encourage the production of a
comprehensive interactive database in which an individual could
possibly belong to overlapping taxonomical groups." "Any
organism can then be characterized by many names because it can
belong to more than one group at once." - Bapteste,
Eric, Yan Boucher. 2008. Lateral gene transfer challenges principles
of microbial systematics. Trends in Microbiology, Vol. 16, No. 5,
pp. 200-207.
Epigenetics
(meaning "above" genetics, as in controlling elements)
Evolutionists
are starting to turn to epigenetics to explain macroevolutionary
changes. However, most of them do not know much about molecular
biology, and they are scrambling to catch up. "During
the twentieth century, evolutionary and molecular biology diverged".
"Few scientists were trained in both fields, and many
biology departments were split up." "Most evolutionary
biologists emphasize... variation". "Inferences
about the historical mechanisms that generate variation are usually
drawn from patterns of association". "The weakness
is that statistical associations are not reliable indicators of
causality." "Claims that are based solely on associations
remain standard in the field." "Evolutionary biologists
will need to be trained in molecular biology". - Dean,
Antony M., Joseph W. Thornton. September 2007. Mechanistic approaches
to the study of evolution: the functional synthesis. Nature Reviews
Genetics, Vol. 8, pp. 675-688.
Here is some of what real molecular biologists have learned, beginning with a March 2008 report in Science News magazine: "Many people regard ribonucleic acid, as RNA is formally known, as 'just a middleman between DNA and protein,' says Claes Wahlestedt, a neuroscientist and genome researcher at the Scripps Research Institute in Jupiter, Fla. Shuttling genetic information from DNA to a cell's protein factories has long been recognized as RNA's day job, summarized" as "DNA makes RNA makes protein." "Some researchers estimate that as much as 98 percent of the human genome is copied into RNA, says Sofie Salama of the University of California, Santa Cruz." "Initial observations of the genome showed islands of protein-coding genes separated by vast oceans of DNA--sometimes called junk DNA--where nothing happened. That would mean that only about 2 percent of the human genome is transcribed into RNA. But recent efforts to map all of the RNA transcripts show that virtually every base pair of DNA in the human genome is copied into at least one RNA molecule."
"More than 20 classes of noncoding RNA have been discovered in the past decade. Many of these RNAs are much smaller than their protein-coding cousins, the messenger RNAs. Some noncoding RNAs contain a mere 20 nucleotides, the chemical units corresponding to letters in the genetic alphabet. Scientists used to throw away such short bits of RNA, thinking the tiny pieces were nothing more than breakdown products of larger molecules--basically garbage, Wahlestedt says."
"Researchers now know that noncoding RNAs get involved in virtually everything that happens in or to a cell, says Georges St. Laurent III, a computational and molecular biologist at George Washington University in Washington, D.C." "They monitor temperature, chemical conditions, electrical currents, and other signals from the environment and then tell the cell how to respond."
"One class of noncoding RNAs, known as microRNAs, modulates production of proteins. MicroRNAs get their name from their minuscule size--most are only about 22 nucleotides long. These short pieces of RNA find and bind to complementary sequences in messenger RNAs. Usually that binding causes the ribosome, the protein-building machinery in a cell, to grind to a halt. The ribosome remains paused until other signals allow it to resume making protein or until the RNA message is destroyed." " 'It's not only important that you make a particular protein, but when and where you make it,' Salama says." - Tina Hesman Saey. March 1, 2008. Micromanagers: New classes of RNAs emerge as key players in the brain. Science News, Vol. 173, No. 9, pp. 136-137.
Non-coding RNAs have risen from "junk" to "drivers of complexity".42 "Sequencing the genomes of 85 species has revealed that in any given organism, increasing biological complexity is correlated with an increasing number of non-protein-coding DNA sequences and not, as previously assumed, with an increasing number of protein-coding genes."42 "The sheer number of non-coding RNAs is estimated in the 100s of thousands."15 "It is clear that tens of thousands may operate within a cell".42
"Interference and activation can be caused by the same transcript".4 "A large part of the transcriptional activity in the human genome is derived from repeat sequences".42 "Repeat elements... occupy 40-45% of a typical mammalian genome".4 "Alu repetitive elements constitute 10% of the human genome".31 "Repeat elements, such as the Alu family in humans and B2 in mice have provided regulatory signals for RNA PolII transcription."7 "Some of the Alu elements... may have functions in stress response, chromatin organization or signaling events in the early embryo. Alu transcripts are... activated by heat shock and DNA damaging agents".42
There are levels of non-coding RNA regulation that have yet to be discovered.42 Studying the old "junk" transcripts can lead to understanding hidden layers of cell regulation and how deregulation can lead to the understanding of human disease.42 "The scientific community is getting more aware of the importance of the formerly abandoned 'junk' DNA. What we have learned so far is likely just the tip of the iceberg."42
It is clear that biological complexity depends less on gene number and more on how those genes are used. Researchers are realizing that regulation is on multiple levels32; there are intricate feedback loops.6 Stretches of DNA can be inactivated by attaching methyl groups. Tiny embryos need to grow according to a body plan organized in steps that have to happen at the right time in the right sequence. Their cells use timers and spatial signals to guide their growth. For example, a signal chemical is made at one end of an embryo and spreads out. Cells act according to how much signal chemical reaches them. Signal chemicals spreading from opposite ends of an embryo can interact to coordinate construction.28
In small genomes, such as yeast, the parts of DNA that regulate a gene are next to the gene. In more complex genomes, such as human and mouse, they can be far apart. Cells have ways, still unknown, of moving sections of chromosomes next to each other to get the right parts together to control gene expression.11 This happens constantly.
To respond to a rapidly changing environment, a creature's genes have to be turned on and off in a highly coordinated way. The genetic network must be stable under a broad range of conditions, but flexible enough to recognize and respond to important signals when things around it change. This operating at the brink of order and chaos is well known to systems scientists. They call such systems critical. This property has now been recognized in plants, animals, and microbes. It allows them to quickly detect and respond to external stimuli, small or large.4
In another surprise to evolutionists,
genes they have long called vestigial junk have a clear purpose.
Some genes are not transcribed into proteins, yet they seem
related to working genes. So evolutionists figured these were
leftover copying mistakes, called them pseudogenes (fake genes),
and ignored them. Researchers studying cancer finally took
a look at a pseudogene in action, and found that it competed with
its working gene for the same non-coding regulatory RNA. Thus
the gene and pseudogene act as decoys for one another, and affect
the regulation of other transcripts. The pseudogene "is not a non-functional relic, but
a modulator of gene expression." This
"could have major implications for understanding mechanisms
of disease, and of cancer in particular." "The authors
find similar associations between other well-known cancer-associated
genes and their corresponding pseudogenes." -
Poliseno, Laura, Leonardo Salmena, Jiangwen Zhang,
Brett Carver, William J. Haveman, Pier Paolo Pandolfi. 24 June 2010. A coding-independent function of gene and pseudogene mRNAs regulates tumour
biology. Nature, Vol. 465, pp. 1033-1038.
Rigoutsos, Isidore, Frank Furnari. 24 June 2010. Decoy for microRNAs.
News & Views, Nature, Vol. 465, pp. 1016-1017.
The Mind of
the Evolutionist
" 'Contemporary evolutionary
thinking maintains that smaller island mammals will rapidly grow
larger towards the optimal size, while bigger animals will rapidly
shrink.' " Evolutionists call this Optimal Body Size
Theory, or OST. A member of a research team was interviewed
about their study that "looked at a theoretical optimum body
size towards which mammals are expected to grow, on both island
communities and on the mainland." " 'There is a
tendency to believe that big animals become very small on islands,
and small animals become very big, due to limited resources or lack
of competition. I've shown that this is just not true, at
least not as a general rule.' " "Incorporating large
data sets that compared body sizes on various islands and on mainland
communities, Dr. Meiri and his colleagues found no such tendency
for bizarrely-sized animals to develop on islands. 'We concluded
that the evolution of body sizes is as random with respect to "isolation"
as on the rest of the planet. This means that you can expect
to find the same sort of patterns on islands and on the mainland.'
Dr. Meiri attributes our widely held misperceptions about
'dragons and dwarfs' to the fact that people tend to notice the
extremes more if they are found on islands." "Darwin's
fascination with the Galapagos island chain... is just one example."
" 'I think it's purely a psychological bias,' Dr. Meiri
concludes. 'It's just magical thinking. Nothing more.'
" - 'Magical Thinking' About Islands an Illusion? Biologist
Refutes Conventional Thinking on Evolution. July 8, 2010. Science
Daily, online news release.
"We found no support for any of the predictions of the optimal size theory." "The concept of a single optimal body size is not supported by the data that were thought most likely to show it." "It is remarkable that this theory fails to apply under the circumstances which best match its predictions (on islands)." - Raia, Pasquale, Francesco Carotenuto, Shai Meiri. 2010. One size does not fit all: no evidence for an optimal body size on islands. Global Ecology and Biogeography, Vol. 19, pp. 475-484.
Evolutionists claim to rely only on natural forces, but natural forces cannot design and build new plants and animals. So they add magical thinking, the perfect description of the evolutionist mind.
Zombie science
"Although
the classical ideal is that scientific theories are evaluated by
a careful teasing-out of their internal logic and external implications,
and checking whether these deductions and predictions are in-line-with
old and new observations; the fact that so many vague, dumb or incoherent
scientific theories are apparently believed by so many scientists
for so many years is suggestive that this ideal does not necessarily
reflect real world practice. In the real world it looks more
like most scientists are quite willing to pursue wrong ideas for
so long as they are rewarded with a better chance of achieving more
grants, publications and status."
To say "that the theory is phoney, and always was phoney, and this is why it so singularly fails to predict reality is regarded as simplistic, crass, merely a sign of lack of sophistication. And anyway, there are... the reputations of numerous scientists who are now successful and powerful on the back of the phoney theory, and who by now control the peer review process (including allocation of grants, publications and jobs) so there is a powerful disincentive against upsetting the apple cart."
"Zombie science is science that is dead but will not lie down." "Zombie science is supported because it is useful propaganda. Zombie science is deployed in arenas such as political rhetoric, public administration, management, public relations, marketing and the mass media generally. It persuades, it constructs taboos, it buttresses some kind of rhetorical attempt to shape mass opinion. Indeed, zombie science often comes across in the mass media as being more plausible than real science." - Charlton, Bruce G. 2008. Zombie science: A sinister consequence of evaluating scientific theories purely on the basis of enlightened self-interest. Medical Hypotheses, Vol. 71, pp. 327-329.
|
Darwin is liked by evolutionists because he liberated science from the straitjacket of observation and opened the door to storytellers. This gave professional evolutionists job security so they can wander through biology labs as if they belong there. ---
David Coppedge You cant win a scientific debate with a storyteller who thinks his imagination is equivalent to scientific evidence. ---
David Coppedge |
**********
1. Alberts, Bruce, Alexander Johnson, Julian Lewis, Martin Raff, Keith Roberts, Peter Walter. 2008. Molecular Biology of The Cell, 5th edition. Garland Science, New York.
2. Anderson, G. M. 1996. Thermodynamics of Natural Systems. John Wiley & Sons, Toronto.
3. Baker, Monya. 10 February 2011. Genomes in Three Dimensions. Nature, Vol. 470, pp. 289-294.
4. Balleza, Enrique, Elena R. Alvarez-Buylla, Alvaro Chaos, Stuart Kauffman, Ilya Shmulevich, Maximino Aldana. June 2008. Critical Dynamics in Genetic Regulatory Networks: Examples from Four Kingdoms. PLoS ONE, Vol. 3, No. 6, pp. 1-10.
5. Bergman, Jerry. August 2000. Do any vestigial organs exist in humans? Technical Journal, Vol. 14, No. 2, pp. 95-98.
6. Brandman, Onn, Tobias Meyer. 17 October 2008. Feedback Loops Shape Cellular Signals in Space and Time. Science, Vol. 322, No. 5900, pp. 390-395.
7. Carninci, Piero, Jun Yasuda, Yoshihide Hayashizaki. 2008. Multifaceted mammalian transcriptome. Current opinion in Cell Biology, Vol. 20, pp. 274-280.
8. Cemic, Ladislav. 2005. Thermodynamics in Mineral Sciences. Springer, New York.
9. Coates, Michael I. 26 March 2009. Beyond the Age of Fishes. Nature, Vol. 458. pp. 413-414.
10. Dagan T., W. Martin. 2006. The tree of one percent. Genome Biology, Vol. 7, No. 10, pp. 118.
11. Dekker, Job. 28 March 2008. Gene Regulation in the Third Dimension. Science, Vol. 319, pp. 1793-1794.
12. Doolittle, W. Ford, Eric Bapteste. February 13, 2007. Pattern pluralism and the Tree of Life hypothesis. Proceedings of the National Academy of Sciences of the United States of America, Vol. 104, No. 7, pp. 2043-2049.
13. Doolittle, W. Ford. 25 June 1999. Phylogenetic Classification and the Universal Tree. Science, Vol. 284, No. 5423, pp. 2124-2128.
14. Duke University Medical Center. "Evolution Of The Human Appendix: A Biological 'Remnant' No More." ScienceDaily 21 August 2009. Retrieved 12 September 2009 <http://www.sciencedaily.com /releases/2009/08/090820175901.htm>.
15. Fahraeus, Robin, Marc Blondel. 2008. Editorial: RNA-assisted protein folding. Biotechnology Journal, Vol. 3, pp. 967-969.
16. Franze, Kristian, Jens Grosche, Serguei N. Skatchkov, Stefan Schinkinger, Christian Foja, Detlev Schild, Ortrud Uckermann, Kort Travis, Andreas Reichenbach, Jochen Guck. May 15, 2007. Muller cells are living optical fibers in the vertebrate retina. PNAS Vol. 104, No. 20, pp. 8287-8292. DOI: 10.1073/pnas.0611180104
17. Gillooly, James F., Andrew P. Allen, Geoffrey B. West, James H. Brown. January 4, 2005. The rate of DNA evolution: Effects of body size and temperature on the molecular clock. Proceedings of the National Academy of Sciences of the United States of America, Vol. 102, No. 1, pp. 140-145.
18. Gish, Duane T., PhD. Biochemistry. January 2007. A Few Reasons an Evolutionary Origin of Life Is Impossible. Impact #403, Acts and Facts, Institute for Creation Research.
19. Glass, John I., Nacyra Assad-Garcia, Nina Alperovich, Shibu Yooseph, Matthew R. Lewis, Mahir Maruf, Clyde A. Hutchison III, Hamilton O. Smith, J. Craig Venter. January 10, 2006. Essential genes of a minimal bacterium. Proceedings of the National Academy of Sciences of the United States of America, Vol. 103, No. 2, pp. 425-430.
20. Jacob, Francois. June 10 1977. Evolution and Tinkering. Science, New Series, Vol. 196, Issue 4295, pp. 1161-1166.
21. Khalturin K, Anton-Erxleben F, Sassmann S, Wittlieb J, Hemmrich G, et al. 2008. A novel gene family controls species-specific morphological traits in Hydra. PLoS Biol 6(11): e278. DOI:10.1371/journal.pbio.0060278
22. Koonin, Eugene V. 12 February 2009. Darwinian evolution in the light of genomics. Nucleic Acids Research, Vol. 37, No. 4, pp. 1011-1034.
23. Koonin, Eugene V. 20 August 2007. The Biological Big Bang model for the major transitions in evolution. Biology Direct, Vol. 2:21, pp. 1-17.
24. Labin, A.M., E. N. Ribak. 16 April 2010. Retinal Glial Cells Enhance Human Vision Acuity. Physical Review Letters, Vol. 104, No. 15, pp. 158102-1 to 4. DOI: 10.1103/PhysRevLett.104.158102
25. Labin, Amichai M., Shadi K. Safuri, Erez N. Ribak, Ido Perlman. 8 July 2014. Muller cells separate between wavelengths to improve day vision with minimal effect upon night vision. Nature Communications, Vol. 5, No. 4319, pp. 1-9. DOI: 10.1038/ncomms5319
26. Lawton, Graham. 21 January 2009. Why Darwin was wrong about the tree of life. New Scientist Magazine, issue 2692.
27. Lee, Michael S.Y., James B. Jago, Diego C. Garcia-Bellido, Gregory D. Edgecombe, James G. Gehling, John R. Paterson. 30 June 2011. Modern optics in exceptionally preserved eyes of Early Cambrian arthropods from Australia. Nature, Vol. 474, pp. 631-634.
28. Lewis Julian. 17 October 2008. From Signals to Patterns: Space, Time, and Mathematics in Developmental Biology. Science, Vol. 322, pp. 399-403.
29. Lowe, Craig B., Gill Bejerano, David Haussler. May 8, 2007. Thousands of human mobile element fragments undergo strong purifying selection near developmental genes. Proceedings of the National Academy of Sciences of the United States of America, Vol. 104, No. 19, pp. 8005-8010.
30. Lunyak, Victoria V., Gratien G. Prefontaine, Esperanza Nunez, Thorsten Cramer, Bong-Gun Ju, Kenneth A. Ohgi, Kasey Hutt, Rosa Roy, Angel Garcia-Diaz, Xiaoyan Zhu, Yun Yung, Lluis Montoliu, Christopher K. Glass, Michael G. Rosenfeld. July 13, 2007. Developmentally Regulated Activation of a SINE B2 Repeat as a Domain Boundary in Organogenesis. Science, Vol. 317, No. 5835, pp. 248-251.
31. Makalowski, Wojciech. May 23, 2003. Not Junk After All. Science, Vol. 300, No. 5623, pp. 1246-1247.
32. Makeyev, Eugene V., Tom Maniatis. 28 March 2008. Multilevel Regulation of Gene Expression by MicroRNAs. Science, Vol. 319, pp. 1789-1790.
34. Pace, John K. II, Clement Gilbert, Marlena S. Clark, Cedric Feschotte. November 4, 2008. Repeated horizontal transfer of a DNA transposon in mammals and other tetrapods. Proceedings of the National Academy of Sciences, Vol. 105, No. 44, pp. 17023-17028.
35. Pilcher, Helen. 19 January 2013. Genes from nowhere: Orphans with a surprising story. New Scientist, No. 2900, pp. 38-41.
36. Prigogine, Ilya, Gregoire Nicolis, Agnes Babloyants. 1972. Thermodynamics of Evolution. Physics Today, Vol. 25, No. 11, pp. 23-44.
37. Reichenbach, Andreas, Andreas Bringmann. May 2013. New functions of Muller cells. Glia, Vol. 61, No. 5, pp. 651-678. DOI: 10.1002/glia.22477
38. Richardson, Michael K., James Hanken, Mayoni L. Gooneratne, Claude Pieau, Albert Raynaud, Lynne Selwood, Glenda M. Wright. July 1997. There is no highly conserved embryonic stage in the vertebrates: implications for current theories of evolution and development. Anatomy and Embryology, Vol. 196, No. 2, pp. 91-106.
39. Shu, D.-G., S. Conway Morris, J. Han, Z.-F. Zhang, K. Yasui, P. Janvier, L. Chen, X.-L. Zhang, J.-N. Liu, Y. Li, H.-Q. Liu. 30 January 2003. Head and backbone of the Early Cambrian vertebrate Haikouichthys. Nature, Vol. 421, pp. 526-529.
40. Smith, H.F., R.E. Fisher, M.L. Everett, A.D. Thomas, R. Randal Bollinger, W. Parker. October 2009. Comparative anatomy and phylogenetic distribution of the mammalian cecal appendix. Journal of Evolutionary Biology, Vol. 22, No. 10, pp. 1984-1999. DOI:10.1111/j.1420-9101.2009.01809.x
41. Smith, Heather F., William Parker, Sanet H. Kotze, Michel Laurin. 2013. Multiple independent appearances of the cecal appendix in mammalian evolution and an investigation of related ecological and anatomical factors. Comptes Rendus Palevol, 16 pages. DOI:10.1016/j.crpv.2012.12.001
42. Szell, Marta, Zsuzsanna Bata-Csorgo, Lajos Kemeny. 2008. The enigmatic world of mRNA-like ncRNAs: Their role in human evolution and in human diseases. Seminars in Cancer Biology, Vol. 18, pp. 141-148.
43. van Steensel, Bas, Job Dekker. October 2010. Genomics tools for unraveling chromosome architecture. Nature Biotechnology, Vol. 28, No. 10, pp. 1089-1093.
44. Weaver, Robert F. 2008. Molecular Biology, Fourth Edition. McGraw-Hill, New York. pp. 660-680.
45. Wissler, Lothar, Jurgen Gadau, Daniel F. Simola, Martin Helmkampf, Erich Bornberg-Bauer. January 24, 2013. Mechanisms and dynamics of orphan gene emergence in insect genomes. Genome Biology and Evolution Advance Access. DOI:10.1093/gbe/evt009
46. Wood, Richard D., Michael Mitchell, John Sgouros, Tomas Lindahl. 16 February 2001. Human DNA Repair Genes. Science, Vol. 291, No. 5507, pp. 1284-1289.
47. Wu D-D, Irwin DM, Zhang Y-P. 2011. De Novo Origin of Human Protein-Coding Genes. PLoS Genet 7(11): e1002379. DOI:10.1371/journal.pgen.1002379
48. Zhu, Min, Wenjin Zhao, Liantao Jia, Jing Lu, Tuo Qiao, Qingming Qu. 26 March 2009. The oldest articulated osteichthyan reveals mosaic gnathostome characters. Nature, Vol. 458, pp. 469-474.
2006 -
2026
John Michael Fischer
mike@newgeology.us
from David F. Coppedge's Creation Evolution Headlines at crev.info